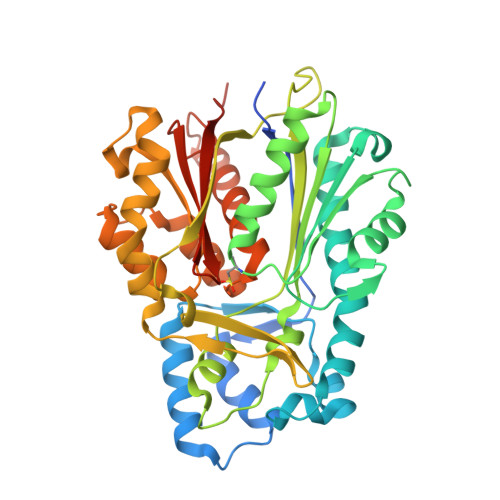
Structural control of polyketide formation in plant-specific polyketide synthases.
Jez, J.M., Austin, M.B., Ferrer, J., Bowman, M.E., Schroder, J., Noel, J.P.(2000) Chem Biol 7: 919-930
- PubMed: 11137815
- DOI: https://doi.org/10.1016/s1074-5521(00)00041-7
- Primary Citation of Related Structures:
1EE0 - PubMed Abstract:
Polyketide synthases (PKSs) generate molecular diversity by utilizing different starter molecules and by controlling the final length of the polyketide. Although exploitation of this mechanistic variability has produced novel polyketides, the structural foundation of this versatility is unclear. Plant-specific PKSs are essential for the biosynthesis of anti-microbial phytoalexins, anthocyanin floral pigments, and inducers of Rhizobium nodulation genes. 2-Pyrone synthase (2-PS) and chalcone synthase (CHS) are plant-specific PKSs that share 74% amino acid sequence identity. 2-PS forms the triketide methylpyrone from an acetyl-CoA starter molecule and two malonyl-CoAs. CHS uses a p-coumaroyl-CoA starter molecule and three malonyl-CoAs to produce the tetraketide chalcone. Our goal was to elucidate the molecular basis of starter molecule selectivity and control of polyketide length in this class of PKS. The 2.05 A resolution crystal structure of 2-PS complexed with the reaction intermediate acetoacetyl-CoA was determined by molecular replacement. 2-PS and CHS share a common three-dimensional fold, a set of conserved catalytic residues, and similar CoA binding sites. However, the active site cavity of 2-PS is smaller than the cavity in CHS. Of the 28 residues lining the 2-PS initiation/elongation cavity, four positions vary in CHS. Point mutations at three of these positions in CHS (T197L, G256L, and S338I) altered product formation. Combining these mutations in a CHS triple mutant (T197L/G256L/S338I) yielded an enzyme that was functionally identical to 2-PS. Structural and functional characterization of 2-PS together with generation of a CHS mutant with an initiation/elongation cavity analogous to 2-PS demonstrates that cavity volume influences the choice of starter molecule and controls the final length of the polyketide. These results provide a structural basis for control of polyketide length in other PKSs, and suggest strategies for further increasing the scope of polyketide biosynthetic diversity.
Organizational Affiliation:
Structural Biology Laboratory, The Salk Institute for Biological Studies, La Jolla, CA 92037, USA.