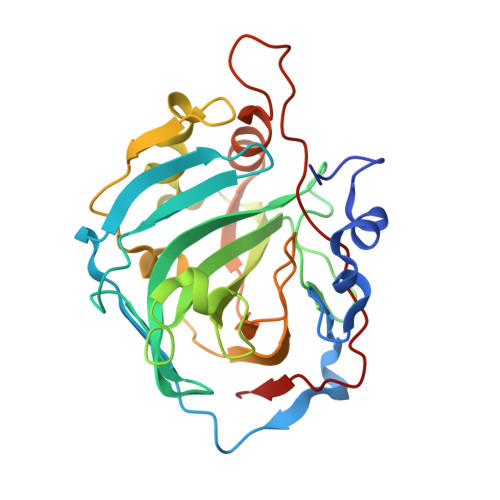
Crystal structure of a zinc-activated variant of human carbonic anhydrase I, CA I Michigan 1: evidence for a second zinc binding site involving arginine coordination.
Ferraroni, M., Tilli, S., Briganti, F., Chegwidden, W.R., Supuran, C.T., Wiebauer, K.E., Tashian, R.E., Scozzafava, A.(2002) Biochemistry 41: 6237-6244
- PubMed: 12009884
- DOI: https://doi.org/10.1021/bi0120446
- Primary Citation of Related Structures:
1J9W, 1JV0 - PubMed Abstract:
The human genetic variant carbonic anhydrase I (CA I) Michigan 1 results from a single point mutation that changes His 67 to Arg in a critical region of the active site. This variant of the zinc metalloenzyme appears to be unique in that it possesses an esterase activity that is specifically enhanced by added free zinc ions. We have determined the three-dimensional structure of human CA I Michigan 1 by X-ray crystallography to a resolution of 2.6 A. In the absence of added zinc ions, the mutated residue, Arg 67, points out of the active site, hydrogen bonding with the carboxylate of Asn 69. This contrasts with the orientation of His 67, in the native isozyme, which points into the active site. The orientations of His 94, His 96, and His 119, that coordinate the catalytic zinc ion, and of the catalytically critical Thr 199-Glu 106 hydrogen bonding system, are largely unchanged in the mutant. The structure of an enzyme adduct with a second zinc bound was determined to a resolution of 2.0 A. The second zinc ion is coordinated to His 64, His 200, and Arg 67. This arginine residue reverses its orientation on zinc binding and turns into the active site. The residues at these three positions have been implicated in determining the specific kinetic properties of native CA I. This is, to our knowledge, the first example of a zinc ion coordinating with an arginine residue in a Zn(II) enzyme.
Organizational Affiliation:
Dipartimento di Chimica, Laboratorio di Chimica Bioinorganica, Università degli Studi di Firenze, Via della Lastruccia, 5, I-50019 Sesto Fiorentino, Italy.