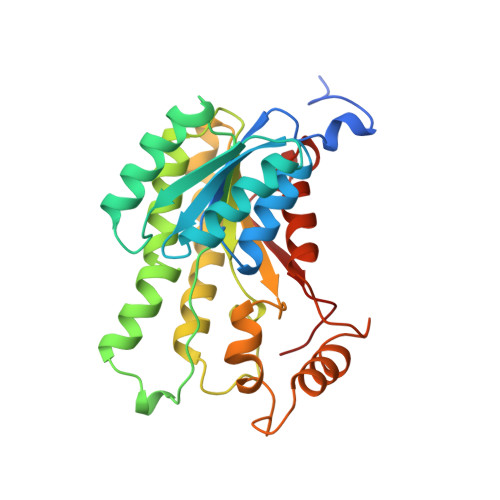
Crystal structure of the ternary complex of 1,3,8-trihydroxynaphthalene reductase from Magnaporthe grisea with NADPH and an active-site inhibitor.
Andersson, A., Jordan, D., Schneider, G., Lindqvist, Y.(1996) Structure 4: 1161-1170
- PubMed: 8939741
- DOI: https://doi.org/10.1016/s0969-2126(96)00124-4
- Primary Citation of Related Structures:
1YBV - PubMed Abstract:
The enzyme 1,3,8-trihydroxynaphthalene reductase (THNR) catalyzes an essential reaction in the biosynthesis of melanin, a black pigment crucial for the pathogenesis of the rice blast fungus, Magnaporthe grisea. The enzyme is the biochemical target of several commercially important fungicides which are used to prevent blast disease in rice plants. We have determined the structure of the ternary complex of THNR with bound NADPH and a fungicide, tricyclazole. Crystallographic analysis showed four identical subunits of THNR to form a tetramer with 222 symmetry. The enzyme subunit consists of a single domain comprising a seven-stranded beta sheet flanked by eight alpha helices; the subunit contains a dinucleotide-binding fold which binds the coenzyme, NADPH. Tricyclazole, an inhibitor of the enzyme, binds at the active site in the vicinity of the NADPH nicotinamide ring. The active site contains a Ser-Tyr-Lys triad which is proposed to participate in catalysis. Coenzyme specificity is partly conferred by the interaction of a single basic residue, Arg39, with the 2' phosphate group of NADPH. The structural model reveals THNR to belong to the family of short chain dehydrogenases. Despite the diversity of the chemical reactions catalyzed by this family of enzymes, their tertiary structures are very similar. In particular THNR has many amino acid sequence identities, and thus most probably high structural similarities, to enzymes involved in fungal aflatoxin synthesis. The structure of THNR in complex with NADPH and tricyclazole provides new insights into the structural basis of inhibitor binding. This new information may aid in the design of new inhibitors for rice crop protection.
Organizational Affiliation:
Department of Molecular Biology, Swedish University of Agricultural Sciences, Uppsala Biomedical Center, Uppsala, Sweden.