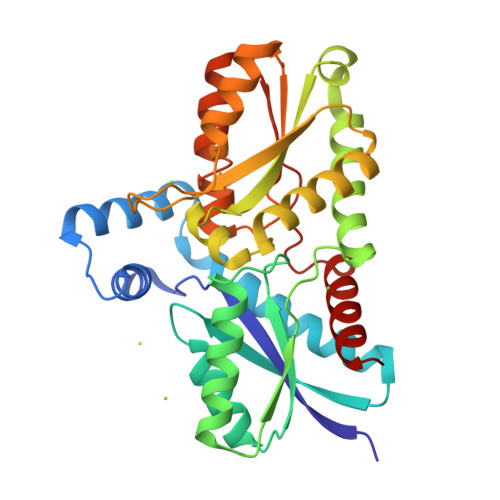
Amino Acid Residues His183 and Glu264 in Bacillus subtilis Ferrochelatase Direct and Facilitate the Insertion of Metal Ion into Protoporphyrin IX
Hansson, M.D., Karlberg, T., Rahardja, M.A., Al-Karadaghi, S., Hansson, M.(2007) Biochemistry 46: 87-94
- PubMed: 17198378
- DOI: https://doi.org/10.1021/bi061760a
- Primary Citation of Related Structures:
2H1V, 2H1W, 2HK6 - PubMed Abstract:
Ferrochelatase catalyzes the terminal step in the heme biosynthetic pathway, i.e., the incorporation of Fe(II) into protoporphyrin IX. Various biochemical and biophysical methods have been used to probe the enzyme for metal binding residues and the location of the active site. However, the location of the metal binding site and the path of the metal into the porphyrin are still disputed. Using site-directed mutagenesis on Bacillus subtilis ferrochelatase we demonstrate that exchange of the conserved residues His183 and Glu264 affects the metal affinity of the enzyme. We also present the first X-ray crystal structure of ferrochelatase with iron. Only a single iron was found in the active site, coordinated in a square pyramidal fashion by two amino acid residues, His183 and Glu264, and three water molecules. This iron was not present in the structure of a His183Ala modified ferrochelatase. The results strongly suggest that the insertion of a metal ion into protoporphyrin IX by ferrochelatase occurs from a metal binding site represented by His183 and Glu264.
Organizational Affiliation:
Department of Biochemistry and Department of Molecular Biophysics, Lund University, Box 124, 221 00 Lund, Sweden. matthias.hansson@biochemistry.lu.se