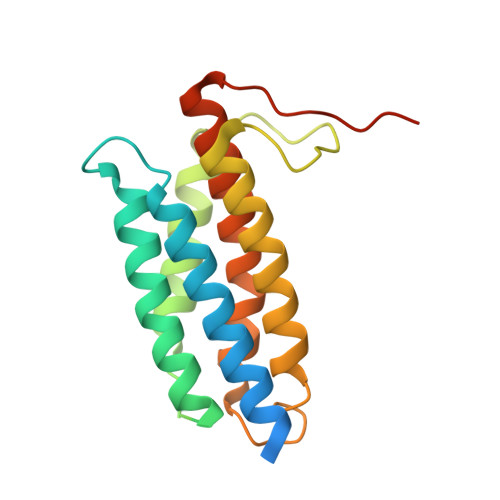
Structure of ATP-Bound Human ATP:Cobalamin Adenosyltransferase.
Schubert, H.L., Hill, C.P.(2006) Biochemistry 45: 15188-15196
- PubMed: 17176040
- DOI: https://doi.org/10.1021/bi061396f
- Primary Citation of Related Structures:
2IDX - PubMed Abstract:
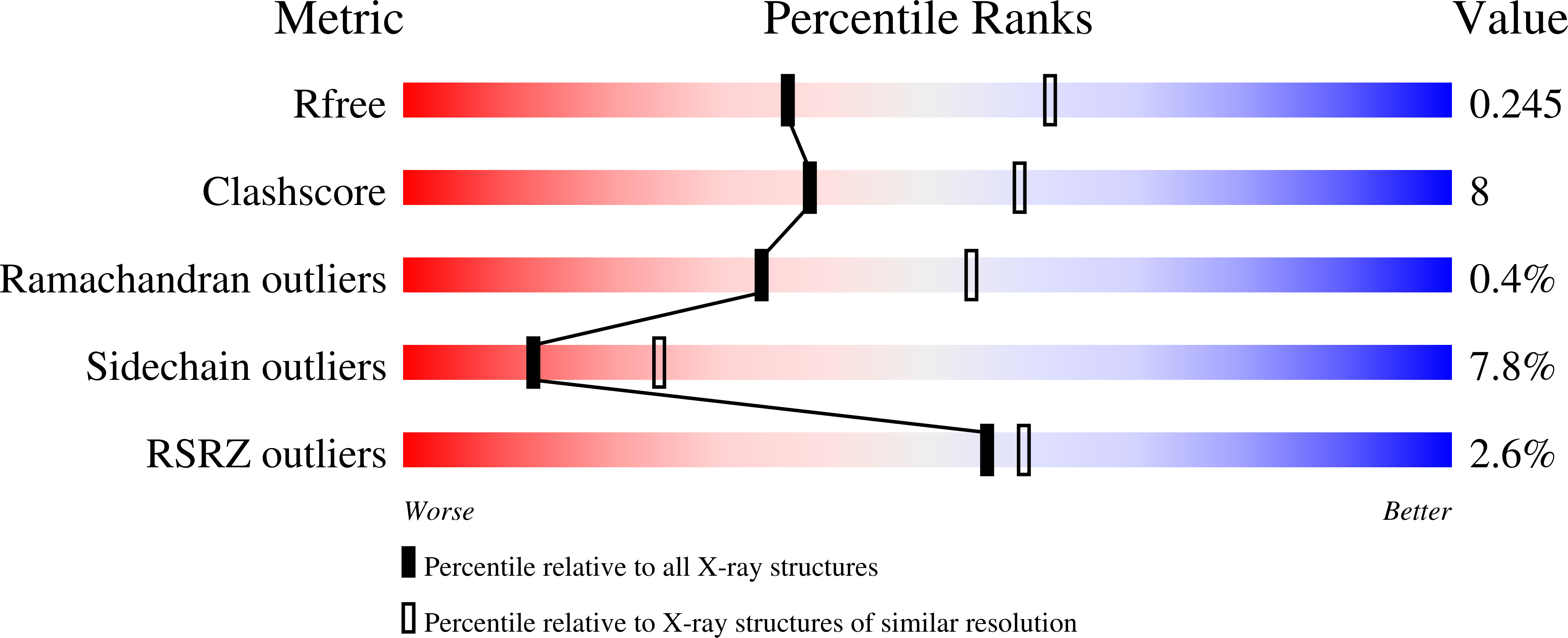
Mutations in the gene encoding human ATP:cobalamin adenosyltransferase (hATR) can result in the metabolic disorder known as methylmalonic aciduria (MMA). This enzyme catalyzes the final step in the conversion of cyanocobalamin (vitamin B12) to the essential human cofactor adenosylcobalamin. Here we present the 2.5 A crystal structure of ATP bound to hATR refined to an Rfree value of 25.2%. The enzyme forms a tightly associated trimer, where the monomer comprises a five-helix bundle and the active sites lie on the subunit interfaces. Only two of the three active sites within the trimer contain the bound ATP substrate, thereby providing examples of apo- and substrate-bound-active sites within the same crystal structure. Comparison of the empty and occupied sites indicates that twenty residues at the enzyme's N-terminus become ordered upon binding of ATP to form a novel ATP-binding site and an extended cleft that likely binds cobalamin. The structure explains the role of 20 invariant residues; six are involved in ATP binding, including Arg190, which hydrogen bonds to ATP atoms on both sides of the scissile bond. Ten of the hydrogen bonds are required for structural stability, and four are in positions to interact with cobalamin. The structure also reveals how the point mutations that cause MMA are deficient in these functions.
Organizational Affiliation:
Department of Biochemistry, University of Utah, Salt Lake City, Utah 84112-5650, USA. heidi@biochem.utah.edu