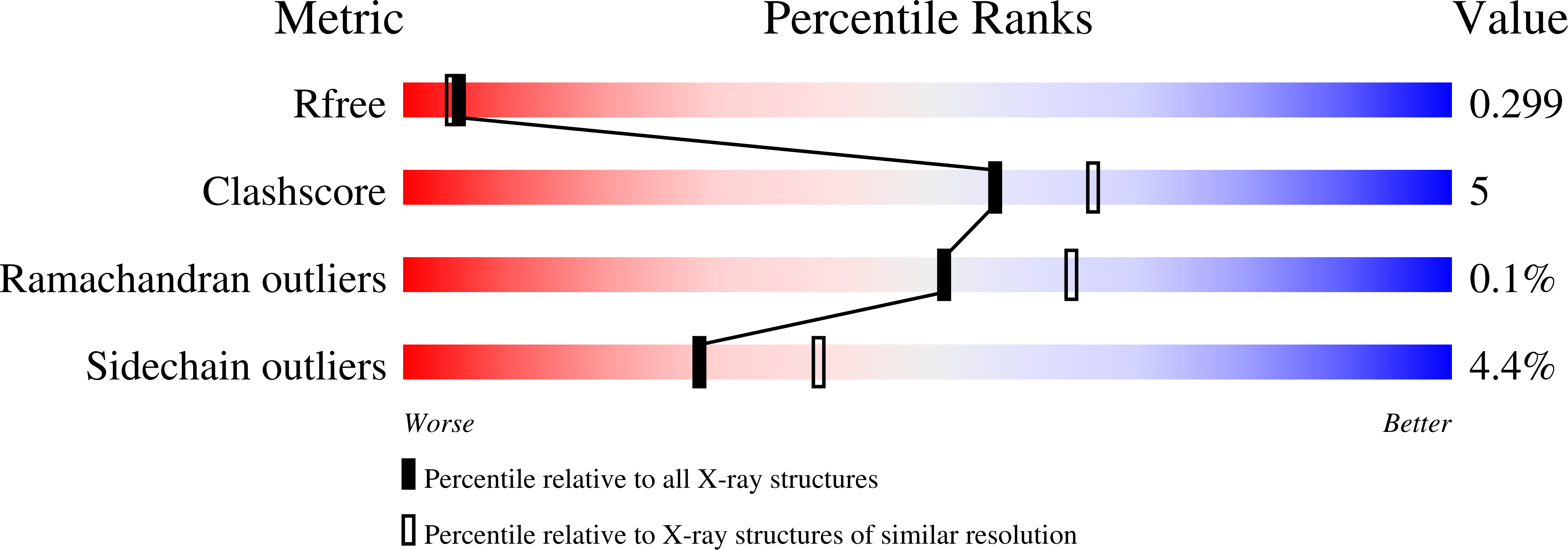
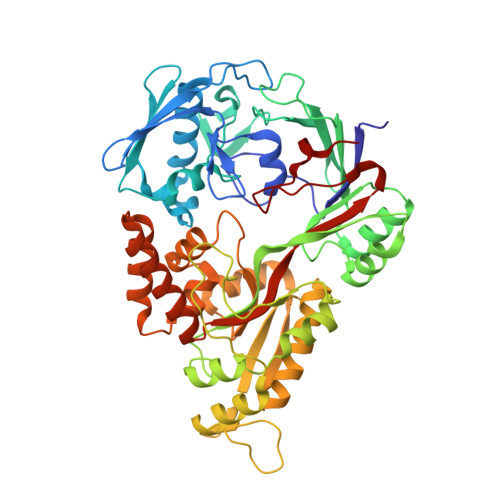
Crystallographic snapshots of the reaction of aromatic C-H with O(2) catalysed by a protein-bound iron complex
Cavazza, C., Bochot, C., Rousselot-Pailley, P., Carpentier, P., Cherrier, M.V., Martin, L., Marchi-Delapierre, C., Fontecilla-Camps, J.C., Menage, S.(2010) Nat Chem 2: 1069-1076
- PubMed: 21107372
- DOI: https://doi.org/10.1038/nchem.841
- Primary Citation of Related Structures:
3MVW, 3MVX, 3MVY, 3MVZ, 3MW0, 3MZ9, 3MZB - PubMed Abstract:
Chemical reactions inside single crystals are quite rare because crystallinity is difficult to retain owing to atomic rearrangements. Protein crystals in general have a high solvent content. This allows for some molecular flexibility, which makes it possible to trap reaction intermediates of enzymatic reactions without disrupting the crystal lattice. A similar approach has not yet been fully implemented in the field of inorganic chemistry. Here, we have combined model chemistry and protein X-ray crystallography to study the intramolecular aromatic dihydroxylation by an arene-containing protein-bound iron complex. The bound complex was able to activate dioxygen in the presence of a reductant, leading to the formation of catechol as the sole product. The structure determination of four of the catalytic cycle intermediates and the end product showed that the hydroxylation reaction implicates an iron peroxo, generated by reductive O(2) activation, an intermediate already observed in iron monooxygenases. This strategy also provided unexpected mechanistic details such as the rearrangement of the iron coordination sphere on metal reduction.
Organizational Affiliation:
Laboratoire de Cristallographie et de Cristallogenèse des Protéines, Institut de Biologie Structurale J.P. Ebel, UMR 5075 CEA, CNRS, Université Joseph Fourier, 41 rue Horowitz, 38027 Grenoble Cedex 1, France. christine.cavazza@ibs.fr