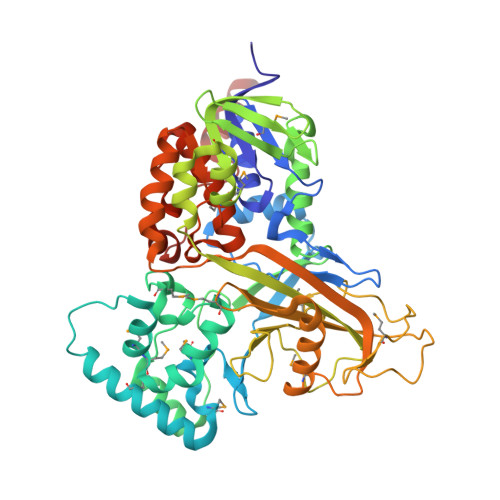
Structural Insight into the Unique Substrate Binding Mechanism and Flavin Redox State of UDP-galactopyranose Mutase from Aspergillus fumigatus.
van Straaten, K.E., Routier, F.H., Sanders, D.A.(2012) J Biological Chem 287: 10780-10790
- PubMed: 22334662
- DOI: https://doi.org/10.1074/jbc.M111.322974
- Primary Citation of Related Structures:
3UKA, 3UKF, 3UKH, 3UKK, 3UKL, 3UKP, 3UKQ - PubMed Abstract:
UDP-galactopyranose mutase (UGM) is a flavin-containing enzyme that catalyzes the reversible conversion of UDP-galactopyranose (UDP-Galp) to UDP-galactofuranose (UDP-Galf). As in prokaryotic UGMs, the flavin needs to be reduced for the enzyme to be active. Here we present the first eukaryotic UGM structures from Aspergillus fumigatus (AfUGM). The structures are of UGM alone, with the substrate UDP-Galp and with the inhibitor UDP. Additionally, we report the structures of AfUGM bound to substrate with oxidized and reduced flavin. These structures provide insight into substrate recognition and structural changes observed upon substrate binding involving the mobile loops and the critical arginine residues Arg-182 and Arg-327. Comparison with prokaryotic UGM reveals that despite low sequence identity with known prokaryotic UGMs the overall fold is largely conserved. Structural differences between prokaryotic UGM and AfUGM result from inserts in AfUGM. A notable difference from prokaryotic UGMs is that AfUGM contains a third flexible loop (loop III) above the si-face of the isoalloxazine ring that changes position depending on the redox state of the flavin cofactor. This loop flipping has not been observed in prokaryotic UGMs. In addition we have determined the crystals structures and steady-state kinetic constants of the reaction catalyzed by mutants R182K, R327K, R182A, and R327A. These results support our hypothesis that Arg-182 and Arg-327 play important roles in stabilizing the position of the diphosphates of the nucleotide sugar and help to facilitate the positioning of the galactose moiety for catalysis.
- Department of Chemistry, University of Saskatchewan, Saskatoon, Saskatchewan S7N 5C9, Canada.
Organizational Affiliation: