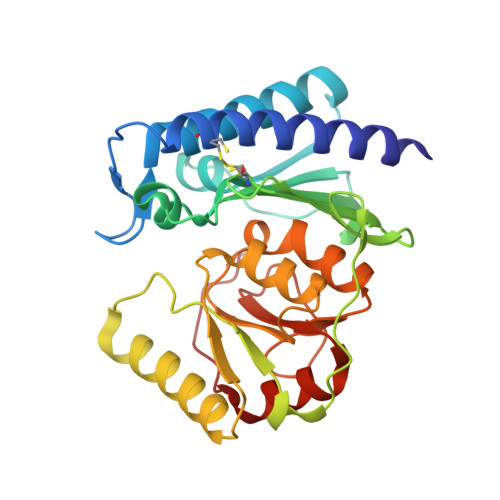
Cloning, Expression, Purification, Crystallization and X-Ray Analysis of Inositol Monophosphatase from Mus Musculus and Homo Sapiens.
Singh, N., Halliday, A.C., Knight, M., Lack, N.A., Lowe, E.D., Churchill, G.C.(2012) Acta Crystallogr Sect F Struct Biol Cryst Commun 68: 1149
- PubMed: 23027737
- DOI: https://doi.org/10.1107/S1744309112035191
- Primary Citation of Related Structures:
4AS4, 4AS5 - PubMed Abstract:
Inositol monophosphatase (IMPase) catalyses the hydrolysis of inositol monophosphate to inositol and is crucial in the phosphatidylinositol (PI) signalling pathway. Lithium, which is the drug of choice for bipolar disorder, inhibits IMPase at therapeutically relevant plasma concentrations. Both mouse IMPase 1 (MmIMPase 1) and human IMPase 1 (HsIMPase 1) were cloned into pRSET5a, expressed in Escherichia coli, purified and crystallized using the sitting-drop method. The structures were solved at resolutions of 2.4 and 1.7 Å, respectively. Comparison of MmIMPase 1 and HsIMPase 1 revealed a core r.m.s. deviation of 0.516 Å.
Organizational Affiliation:
Department of Pharmacology, University of Oxford, Mansfield Road, Oxford OX1 3QT, England.