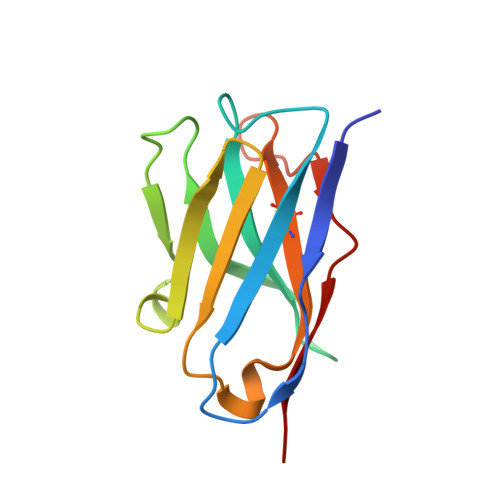
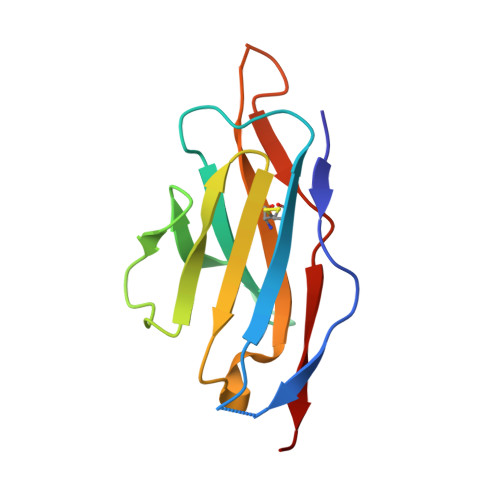
Single-chain antibody-fragment M6P-1 possesses a mannose 6-phosphate monosaccharide-specific binding pocket that distinguishes N-glycan phosphorylation in a branch-specific manner.
Blackler, R.J., Evans, D.W., Smith, D.F., Cummings, R.D., Brooks, C.L., Braulke, T., Liu, X., Evans, S.V., Muller-Loennies, S.(2016) Glycobiology 26: 181-192
- PubMed: 26503547
- DOI: https://doi.org/10.1093/glycob/cwv093
- Primary Citation of Related Structures:
4RZC - PubMed Abstract:
The acquisition of mannose 6-phosphate (Man6P) on N-linked glycans of lysosomal enzymes is a structural requirement for their transport from the Golgi apparatus to lysosomes mediated by the mannose 6-phosphate receptors, 300 kDa cation-independent mannose 6-phosphate receptor (MPR300) and 46 kDa cation-dependent mannose 6-phosphate receptor (MPR46). Here we report that the single-chain variable domain (scFv) M6P-1 is a unique antibody fragment with specificity for Man6P monosaccharide that, through an array-screening approach against a number of phosphorylated N-glycans, is shown to bind mono- and diphosphorylated Man6 and Man7 glycans that contain terminal αMan6P(1 → 2)αMan(1 → 3)αMan. In contrast to MPR300, scFv M6P-1 does not bind phosphodiesters, monophosphorylated Man8 or mono- or diphosphorylated Man9 structures. Single crystal X-ray diffraction analysis to 2.7 Å resolution of Fv M6P-1 in complex with Man6P reveals that specificity and affinity is achieved via multiple hydrogen bonds to the mannose ring and two salt bridges to the phosphate moiety. In common with both MPRs, loss of binding was observed for scFv M6P-1 at pH values below the second pKa of Man6P (pKa = 6.1). The structures of Fv M6P-1 and the MPRs suggest that the change of the ionization state of Man6P is the main driving force for the loss of binding at acidic lysosomal pH (e.g. lysosome pH ∼ 4.6), which provides justification for the evolution of a lysosomal enzyme transport pathway based on Man6P recognition.
Organizational Affiliation:
Department of Biochemistry and Microbiology, University of Victoria, Victoria, BC Canada V8P 3P6.