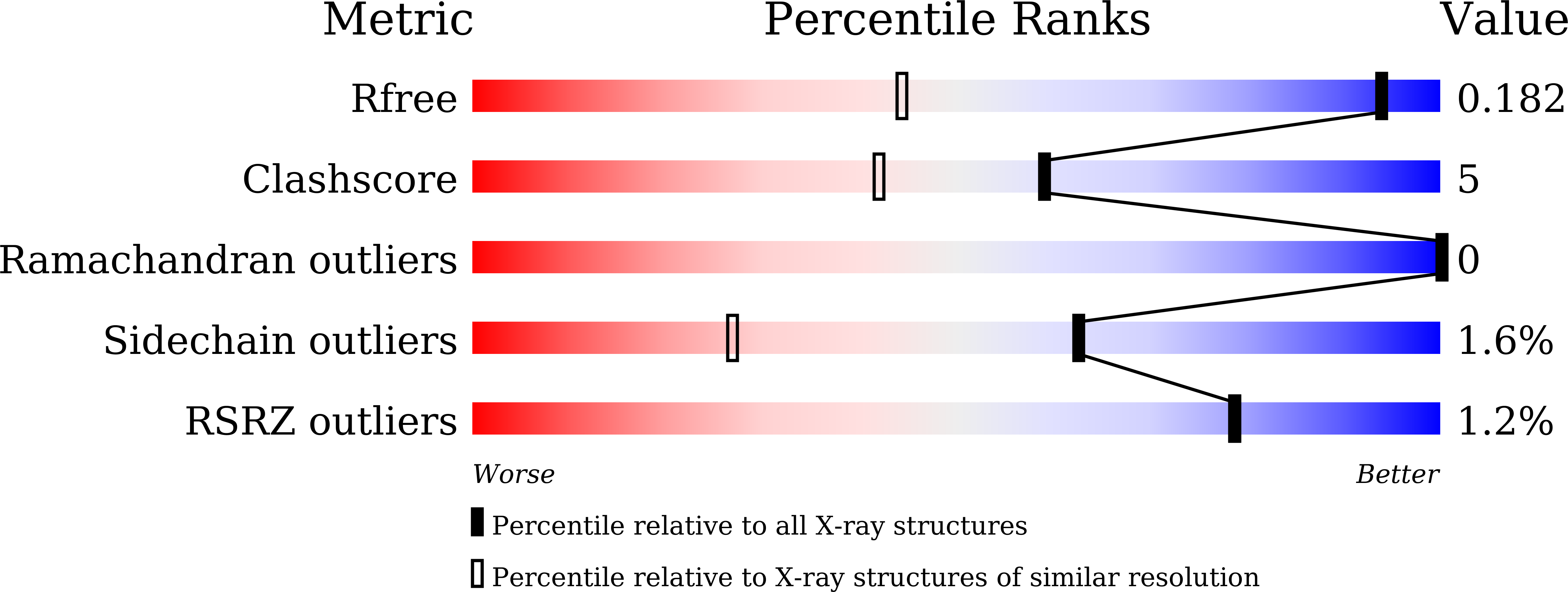
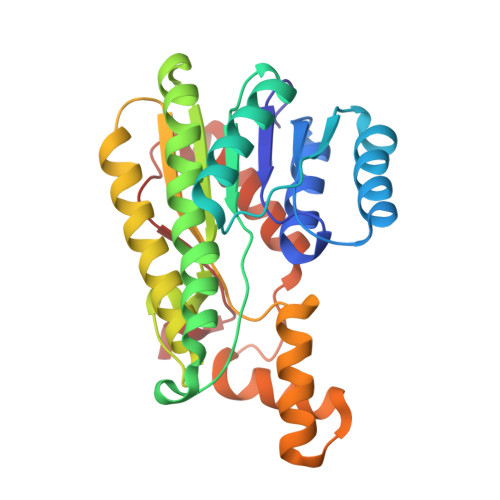
Structural insights into the catalytic reaction trigger and inhibition of D-3-hydroxybutyrate dehydrogenase
Kanazawa, H., Hoque, M.M., Tsunoda, M., Suzuki, K., Yamamoto, T., Kawai, G., Kondo, J., Takenaka, A.(2016) Acta Crystallogr F Struct Biol Commun 72: 507-515
- PubMed: 27380367
- DOI: https://doi.org/10.1107/S2053230X16007767
- Primary Citation of Related Structures:
5B4T, 5B4U, 5B4V - PubMed Abstract:
D-3-Hydroxybutyrate dehydrogenase catalyzes the reversible conversion of acetoacetate and D-3-hydroxybutyrate. These ketone bodies are both energy-storage forms of acetyl-CoA. In order to clarify the structural mechanisms of the catalytic reaction with the cognate substrate D-3-hydroxybutyrate and of the inhibition of the reaction by inhibitors, the enzyme from Alcaligenes faecalis has been analyzed by X-ray crystallography in liganded states with the substrate and with two types of inhibitor: malonate and methylmalonate. In each subunit of the tetrameric enzyme, the substrate is trapped on the nicotinamide plane of the bound NAD(+). An OMIT map definitively shows that the bound ligand is D-3-hydroxybutyrate and not acetoacetate. The two carboxylate O atoms form four hydrogen bonds to four conserved amino-acid residues. The methyl group is accommodated in the nearby hydrophobic pocket so that the formation of a hydrogen bond from the OH group of the substrate to the hydroxy group of Tyr155 at the active centre is facilitated. In this geometry, the H atom attached to the C(3) atom of the substrate in the sp(3) configuration is positioned at a distance of 3.1 Å from the nicotinamide C(4) atom in the direction normal to the plane. In addition, the donor-acceptor relationship of the hydrogen bonds suggests that the Tyr155 OH group is allowed to ionize by the two donations from the Ser142 OH group and the ribose OH group. A comparison of the protein structures with and without ligands indicates that the Gln196 residue of the small movable domain participates in the formation of additional hydrogen bonds. It is likely that this situation can facilitate H-atom movements as the trigger of the catalytic reaction. In the complexes with inhibitors, however, their principal carboxylate groups interact with the enzyme in a similar way, while the interactions of other groups are changed. The crucial determinant for inhibition is that the inhibitors have no active H atom at C(3). A second determinant is the Tyr155 OH group, which is perturbed by the inhibitors to donate its H atom for hydrogen-bond formation, losing its nucleophilicity.
Organizational Affiliation:
Department of Materials and Life Sciences, Sophia University, Kioi-cho, Chiyoda-ku, Tokyo 102-8554, Japan.