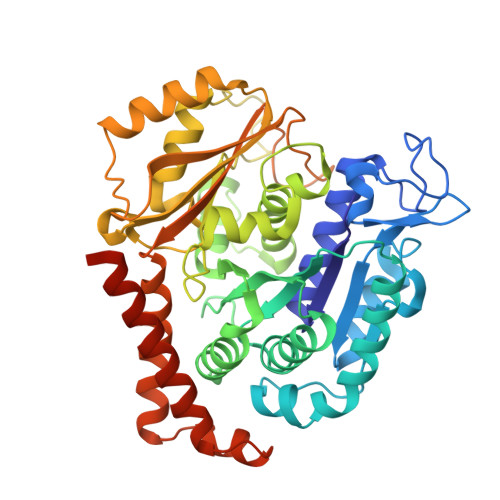
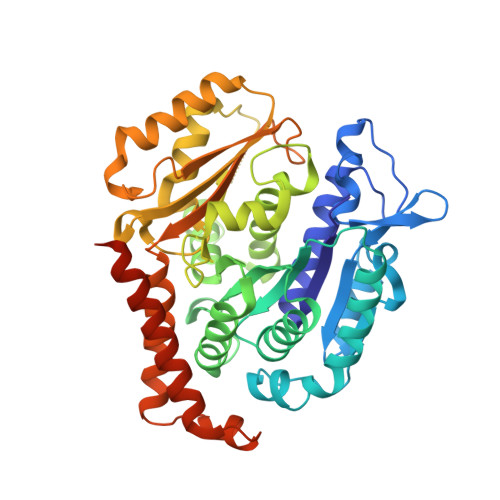
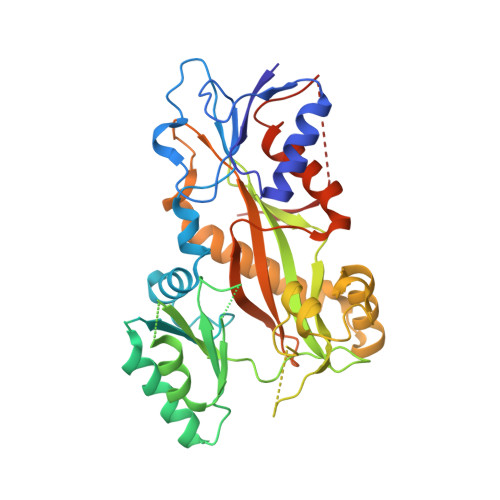
Rational Design of a Novel Tubulin Inhibitor with a Unique Mechanism of Action.
Muhlethaler, T., Milanos, L., Ortega, J.A., Blum, T.B., Gioia, D., Roy, B., Prota, A.E., Cavalli, A., Steinmetz, M.O.(2022) Angew Chem Int Ed Engl 61: e202204052-e202204052
- PubMed: 35404502
- DOI: https://doi.org/10.1002/anie.202204052
- Primary Citation of Related Structures:
5SB3, 5SB4, 5SB5, 5SB6, 5SB7, 7Z7D - PubMed Abstract:
In this study, we capitalized on our previously performed crystallographic fragment screen and developed the antitubulin small molecule Todalam with only two rounds of straightforward chemical synthesis. Todalam binds to a novel tubulin site, disrupts microtubule networks in cells, arrests cells in G2/M, induces cell death, and synergizes with vinblastine. The compound destabilizes microtubules by acting as a molecular plug that sterically inhibits the curved-to-straight conformational switch in the α-tubulin subunit, and by sequestering tubulin dimers into assembly incompetent oligomers. Our results describe for the first time the generation of a fully rationally designed small molecule tubulin inhibitor from a fragment, which displays a unique molecular mechanism of action. They thus demonstrate the usefulness of tubulin-binding fragments as valuable starting points for innovative antitubulin drug and chemical probe discovery campaigns.
Organizational Affiliation:
Laboratory of Biomolecular Research, Department of Biology and Chemistry, Paul Scherrer Institut, 5232, Villigen PSI, Switzerland.