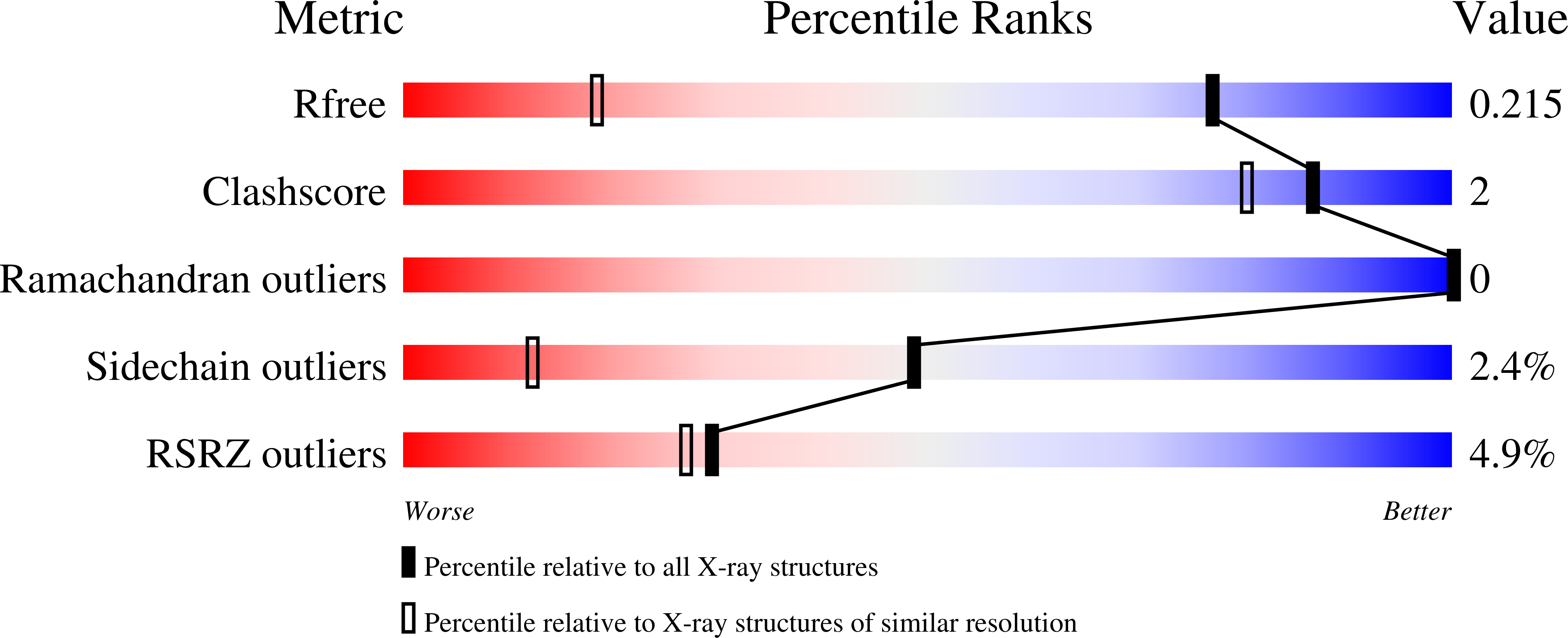
A Unique Approach to Design Potent and Selective Cyclic Adenosine Monophosphate Response Element Binding Protein, Binding Protein (CBP) Inhibitors.
Bronner, S.M., Murray, J., Romero, F.A., Lai, K.W., Tsui, V., Cyr, P., Beresini, M.H., de Leon Boenig, G., Chen, Z., Choo, E.F., Clark, K.R., Crawford, T.D., Jayaram, H., Kaufman, S., Li, R., Li, Y., Liao, J., Liang, X., Liu, W., Ly, J., Maher, J., Wai, J., Wang, F., Zheng, A., Zhu, X., Magnuson, S.(2017) J Med Chem 60: 10151-10171
- PubMed: 29155580
- DOI: https://doi.org/10.1021/acs.jmedchem.7b01372
- Primary Citation of Related Structures:
5W0E, 6AXQ, 6AY3, 6AY5 - PubMed Abstract:
The epigenetic regulator CBP/P300 presents a novel therapeutic target for oncology. Previously, we disclosed the development of potent and selective CBP bromodomain inhibitors by first identifying pharmacophores that bind the KAc region and then building into the LPF shelf. Herein, we report the "hybridization" of a variety of KAc-binding fragments with a tetrahydroquinoline scaffold that makes optimal interactions with the LPF shelf, imparting enhanced potency and selectivity to the hybridized ligand. To demonstrate the utility of our hybridization approach, two analogues containing unique Asn binders and the optimized tetrahydroquinoline moiety were rapidly optimized to yield single-digit nanomolar inhibitors of CBP with exquisite selectivity over BRD4(1) and the broader bromodomain family.
Organizational Affiliation:
Genentech, Inc. , 1 DNA Way, South San Francisco, California 94080, United States.