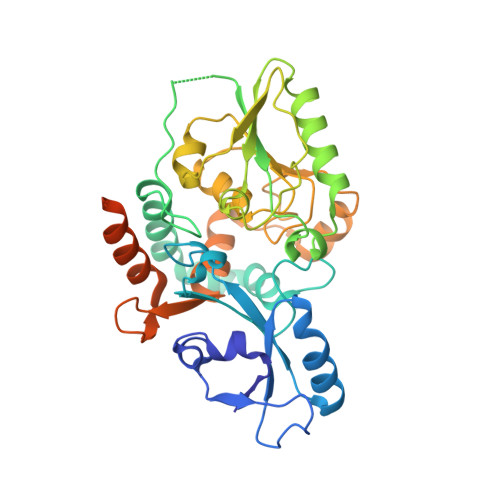
Biosynthesis of a conserved glycolipid anchor for Gram-negative bacterial capsules.
Doyle, L., Ovchinnikova, O.G., Myler, K., Mallette, E., Huang, B.S., Lowary, T.L., Kimber, M.S., Whitfield, C.(2019) Nat Chem Biol 15: 632-640
- PubMed: 31036922
- DOI: https://doi.org/10.1038/s41589-019-0276-8
- Primary Citation of Related Structures:
6MGB, 6MGC, 6MGD - PubMed Abstract:
Several important Gram-negative bacterial pathogens possess surface capsular layers composed of hypervariable long-chain polysaccharides linked via a conserved 3-deoxy-β-D-manno-oct-2-ulosonic acid (β-Kdo) oligosaccharide to a phosphatidylglycerol residue. The pathway for synthesis of the terminal glycolipid was elucidated by determining the structures of reaction intermediates. In Escherichia coli, KpsS transfers a single Kdo residue to phosphatidylglycerol; this primer is extended using a single enzyme (KpsC), possessing two cytidine 5'-monophosphate (CMP)-Kdo-dependent glycosyltransferase catalytic centers with different linkage specificities. The structure of the N-terminal β-(2→4) Kdo transferase from KpsC reveals two α/β domains, supplemented by several helices. The N-terminal Rossmann-like domain, typically responsible for acceptor binding, is severely reduced in size compared with canonical GT-B folds in glycosyltransferases. The similar structure of the C-terminal β-(2→7) Kdo transferase indicates a past gene duplication event. Both Kdo transferases have a narrow active site tunnel, lined with key residues shared with GT99 β-Kdo transferases. This enzyme provides the prototype for the GT107 family.
Organizational Affiliation:
Department of Molecular and Cellular Biology, University of Guelph, Guelph, Ontario, Canada.