From tryptophan-based amides to tertiary amines: Optimization of a butyrylcholinesterase inhibitor series.
Meden, A., Knez, D., Brazzolotto, X., Nachon, F., Dias, J., Svete, J., Stojan, J., Groselj, U., Gobec, S.(2022) Eur J Med Chem 234: 114248-114248
- PubMed: 35299116
- DOI: https://doi.org/10.1016/j.ejmech.2022.114248
- Primary Citation of Related Structures:
7BO3, 7BO4 - PubMed Abstract:
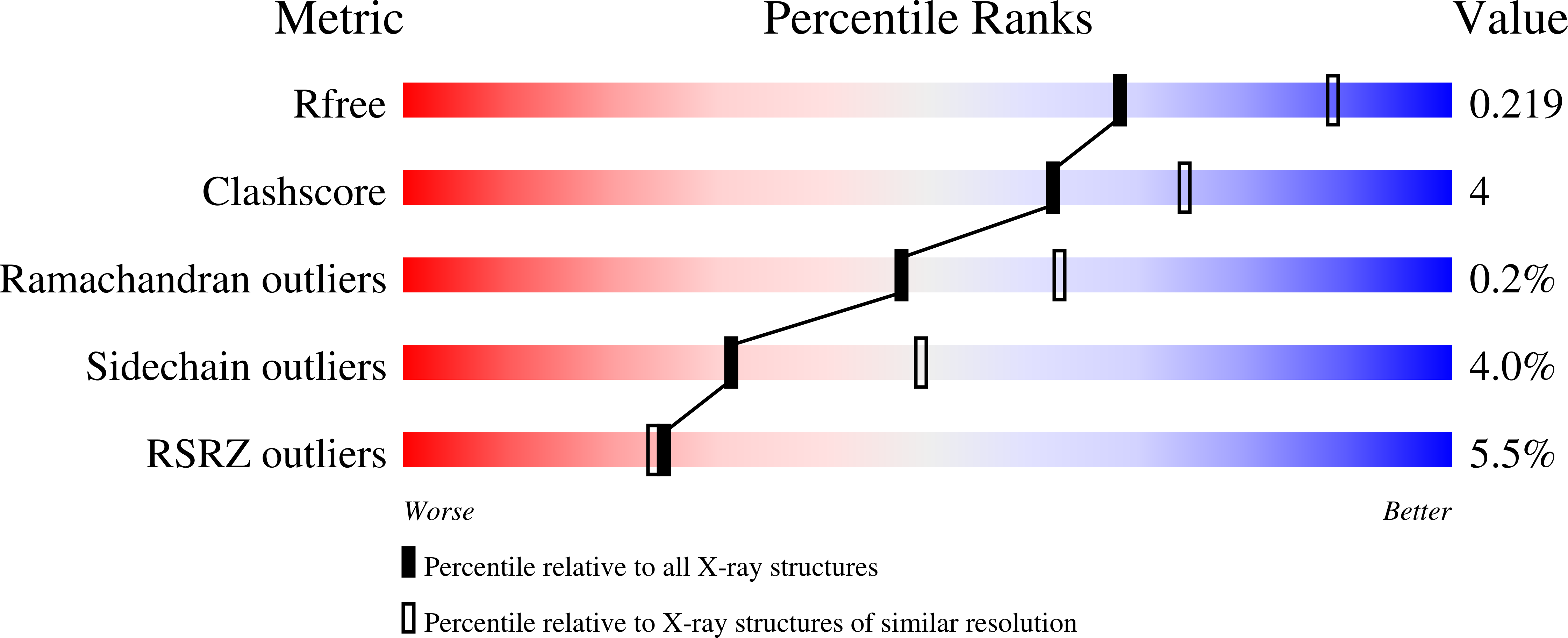
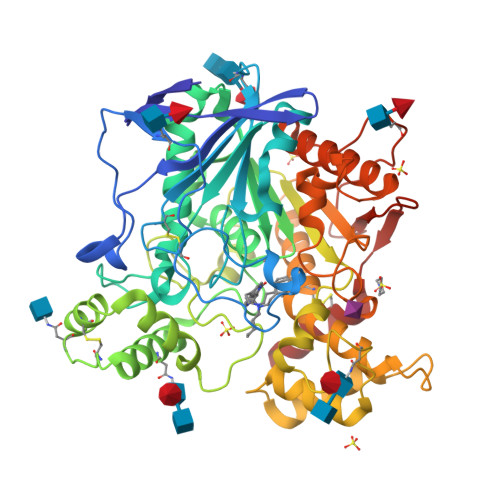
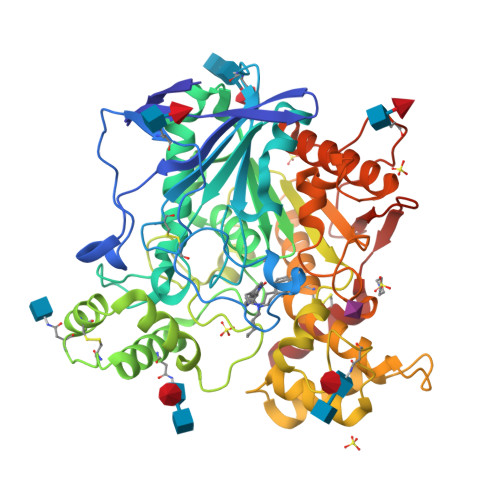
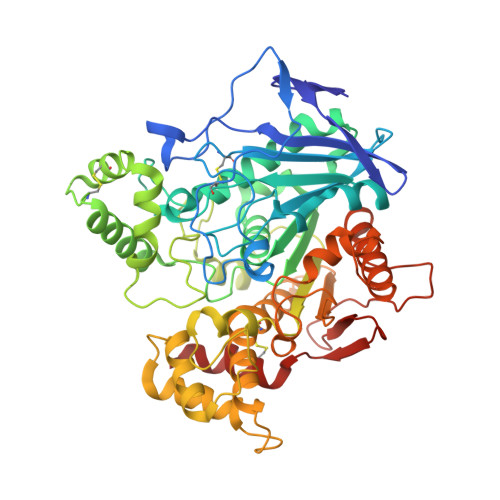
Lead optimization of a series of tryptophan-based nanomolar butyrylcholinesterase (BChE) inhibitors led to tertiary amines as highly potent, achiral, sp 3 -rich analogues with better synthetic accessibility and high selectivity over acetylcholinesterase (one to ten thousandfold). Taking it one step further, the introduction of a carbamate warhead on the well-explored reversible scaffold allowed conversion to pseudoirreversible inhibitors that bound covalently to BChE and prolonged the duration of inhibition (half-life of 14.8 h for compound 45a-carbamoylated enzyme). Additionally, N-hydroxyindole was discovered as a novel leaving group chemotype. The covalent mechanism of action was confirmed by time-dependency experiments, progress curve analysis, and indirectly by co-crystallization with the human recombinant enzyme. Two crystal structures of BChE-inhibitor complexes were solved and coupled with the supporting molecular dynamics simulations increased our understanding of the structure-activity relationship, while also providing the necessary structural information for future optimization of this series. Overall, this research demonstates the high versatility and potential of this series of BChE inhibitors.
Organizational Affiliation:
University of Ljubljana, Faculty of Pharmacy, Aškerčeva 7, SI-1000, Ljubljana, Slovenia.