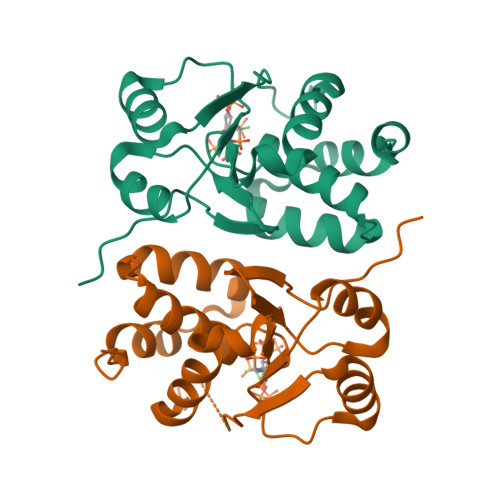
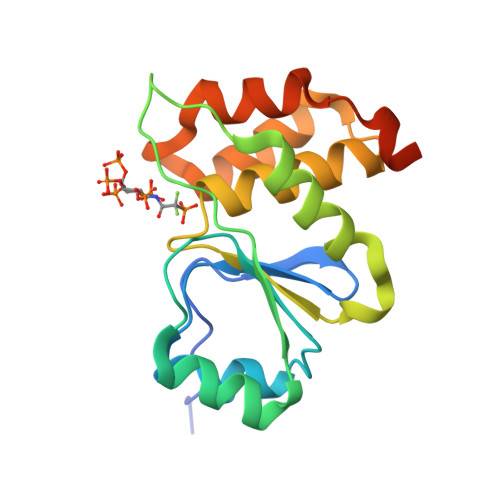
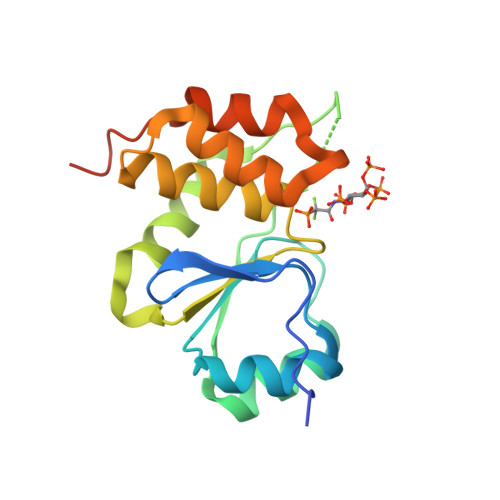
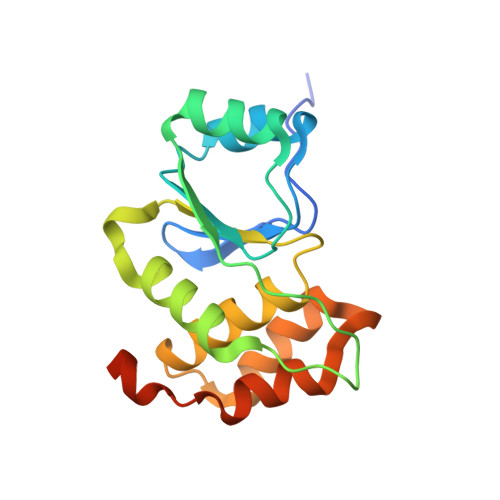
A structural expose of noncanonical molecular reactivity within the protein tyrosine phosphatase WPD loop.
Wang, H., Perera, L., Jork, N., Zong, G., Riley, A.M., Potter, B.V.L., Jessen, H.J., Shears, S.B.(2022) Nat Commun 13: 2231-2231
- PubMed: 35468885
- DOI: https://doi.org/10.1038/s41467-022-29673-y
- Primary Citation of Related Structures:
7MOD, 7MOE, 7MOF, 7MOG, 7MOH, 7MOI, 7MOJ, 7MOK, 7MOL, 7MOM - PubMed Abstract:
Structural snapshots of protein/ligand complexes are a prerequisite for gaining atomic level insight into enzymatic reaction mechanisms. An important group of enzymes has been deprived of this analytical privilege: members of the protein tyrosine phosphatase (PTP) superfamily with catalytic WPD-loops lacking the indispensable general-acid/base within a tryptophan-proline-aspartate/glutamate context. Here, we provide the ligand/enzyme crystal complexes for one such PTP outlier: Arabidopsis thaliana Plant and Fungi Atypical Dual Specificity Phosphatase 1 (AtPFA-DSP1), herein unveiled as a regioselective and efficient phosphatase towards inositol pyrophosphate (PP-InsP) signaling molecules. Although the WPD loop is missing its canonical tripeptide motif, this structural element contributes to catalysis by assisting PP-InsP delivery into the catalytic pocket, for a choreographed exchange with phosphate reaction product. Subsequently, an intramolecular proton donation by PP-InsP substrate is posited to substitute functionally for the absent aspartate/glutamate general-acid. Overall, we expand mechanistic insight into adaptability of the conserved PTP structural elements.
Organizational Affiliation:
Signal Transduction Laboratory, National Institute of Environmental Health Sciences, National Institutes of Health, Research Triangle Park, NC, 27709, USA. huanchen.wang@nih.gov.