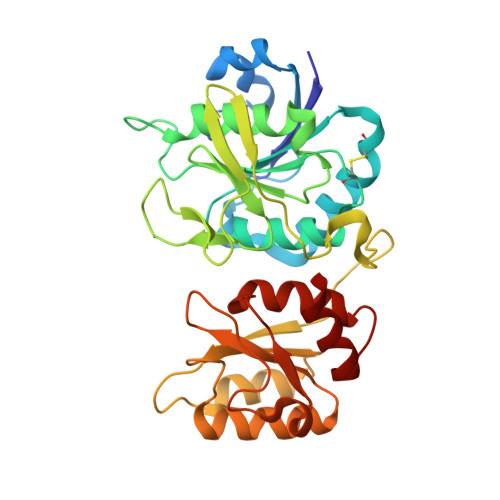
Structural Aspects of E. coli Type II Asparaginase in Complex with Its Secondary Product L-Glutamate.
Maggi, M., Scotti, C.(2022) Int J Mol Sci 23
- PubMed: 35682622
- DOI: https://doi.org/10.3390/ijms23115942
- Primary Citation of Related Structures:
7R57, 7R5Q - PubMed Abstract:
Bacterial L-asparaginases are amidohydrolases (EC 3.5.1.1) capable of deaminating L-asparagine and, with reduced efficiency, L-glutamine. Interest in the study of L-asparaginases is driven by their use as biodrugs for the treatment of acute lymphoblastic leukemia. Here, we report for the first time the description of the molecular structure of type II asparaginase from Escherichia coli in complex with its secondary product, L-glutamate. To obtain high-quality crystals, we took advantage of the N24S variant, which has structural and functional features similar to the wild-type enzyme, but improved stability, and which yields more ordered crystals. Analysis of the structure of the N24S-L-glutamate complex (N24S-GLU) and comparison with its apo and L-aspartate-bound form confirmed that the enzyme-reduced catalytic efficiency in the presence of L-glutamine is due to L-glutamine misfitting into the enzyme-binding pocket, which causes a local change in the catalytic center geometry. Moreover, a tight interaction between the two protomers that form the enzyme active site limits the capability of L-glutamine to fit into (and to exit from) the binding pocket of E. coli L-asparaginase, explaining why the enzyme has lower glutaminolytic activity compared to other enzymes of the same family, in particular the Erwinia chrysanthemi one.
Organizational Affiliation:
Unit of Immunology and General Pathology, Department of Molecular Medicine, University of Pavia, 27100 Pavia, Italy.