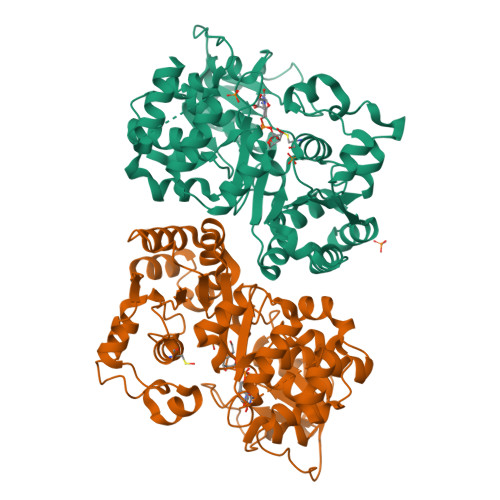
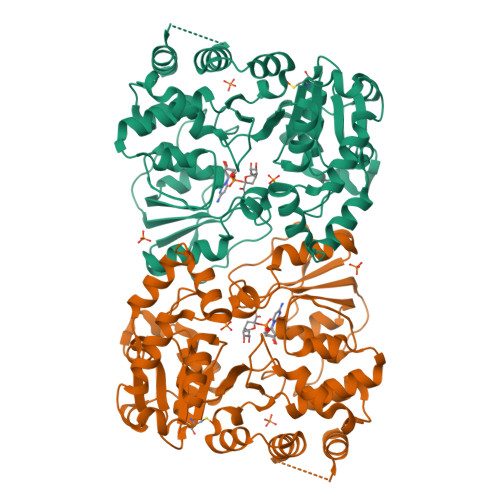
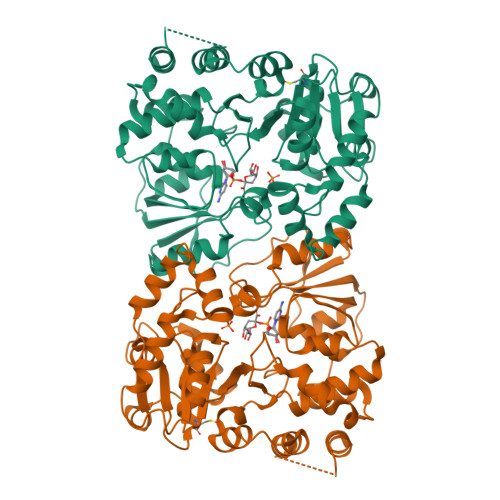
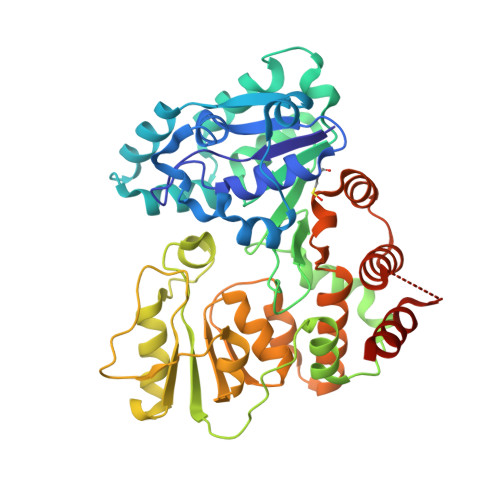
The retaining beta-Kdo glycosyltransferase WbbB uses a double-displacement mechanism with an intermediate adduct rearrangement step.
Forrester, T.J.B., Ovchinnikova, O.G., Li, Z., Kitova, E.N., Nothof, J.T., Koizumi, A., Klassen, J.S., Lowary, T.L., Whitfield, C., Kimber, M.S.(2022) Nat Commun 13: 6277-6277
- PubMed: 36271007
- DOI: https://doi.org/10.1038/s41467-022-33988-1
- Primary Citation of Related Structures:
8CSB, 8CSC, 8CSD, 8CSE, 8CSF - PubMed Abstract:
WbbB, a lipopolysaccharide O-antigen synthesis enzyme from Raoultella terrigena, contains an N-terminal glycosyltransferase domain with a highly modified architecture that adds a terminal β-Kdo (3-deoxy-D-manno-oct-2-ulosonic acid) residue to the O-antigen saccharide, with retention of stereochemistry. We show, using mass spectrometry, that WbbB forms a covalent adduct between the catalytic nucleophile, Asp232, and Kdo. We also determine X-ray structures for the CMP-β-Kdo donor complex, for Kdo-adducts with D232N and D232C WbbB variants, for a synthetic disaccharide acceptor complex, and for a ternary complex with both a Kdo-adduct and the acceptor. Together, these structures show that the enzyme-linked Asp232-Kdo adduct rotates to reposition the Kdo into a second sub-site, which then transfers Kdo to the acceptor. Retaining glycosyltransferases were thought to use only the front-side S N i substitution mechanism; here we show that retaining glycosyltransferases can also potentially use double-displacement mechanisms, but incorporating an additional catalytic subsite requires rearrangement of the protein's architecture.
Organizational Affiliation:
Department of Molecular and Cellular Biology, University of Guelph, 50 Stone Road E., Guelph, ON, N1G 2W1, Canada.