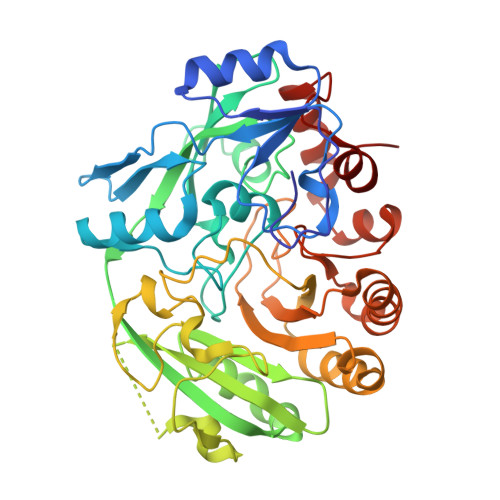
Discovery and biochemical characterization of thermostable glycerol oxidases.
Santema, L.L., Rotilio, L., Xiang, R., Tjallinks, G., Guallar, V., Mattevi, A., Fraaije, M.W.(2024) Appl Microbiol Biotechnol 108: 61-61
- PubMed: 38183484
- DOI: https://doi.org/10.1007/s00253-023-12883-9
- Primary Citation of Related Structures:
8OT8 - PubMed Abstract:
Alditol oxidases are promising tools for the biocatalytic oxidation of glycerol to more valuable chemicals. By integrating in silico bioprospecting with cell-free protein synthesis and activity screening, an effective pipeline was developed to rapidly identify enzymes that are active on glycerol. Three thermostable alditol oxidases from Actinobacteria Bacterium, Streptomyces thermoviolaceus, and Thermostaphylospora chromogena active on glycerol were discovered. The characterization of these three flavoenzymes demonstrated their glycerol oxidation activities, preference for alkaline conditions, and excellent thermostabilities with melting temperatures higher than 75 °C. Structural elucidation of the alditol oxidase from Actinobacteria Bacterium highlighted a constellation of side chains that engage the substrate through several hydrogen bonds, a histidine residue covalently bound to the FAD prosthetic group, and a tunnel leading to the active site. Upon computational simulations of substrate binding, a double mutant targeting a residue pair at the tunnel entrance was created and found to display an improved thermal stability and catalytic efficiency for glycerol oxidation. The hereby described alditol oxidases form a valuable panel of oxidative biocatalysts that can perform regioselective oxidation of glycerol and other polyols. KEY POINTS: • Rapid pipeline designed to identify putative oxidases • Biochemical and structural characterization of alditol oxidases • Glycerol oxidation to more valuable derivatives.
Organizational Affiliation:
Molecular Enzymology, University of Groningen, Nijenborgh 4, 9747AG, Groningen, The Netherlands.