Structural characterization of the active site of the PduO-type ATP:Co(I)rrinoid adenosyltransferase from Lactobacillus reuteri.
St Maurice, M., Mera, P.E., Taranto, M.P., Sesma, F., Escalante-Semerena, J.C., Rayment, I.(2007) J Biol Chem 282: 2596-2605
- PubMed: 17121823
- DOI: https://doi.org/10.1074/jbc.M609557200
- Primary Citation of Related Structures:
2NT8 - PubMed Abstract:
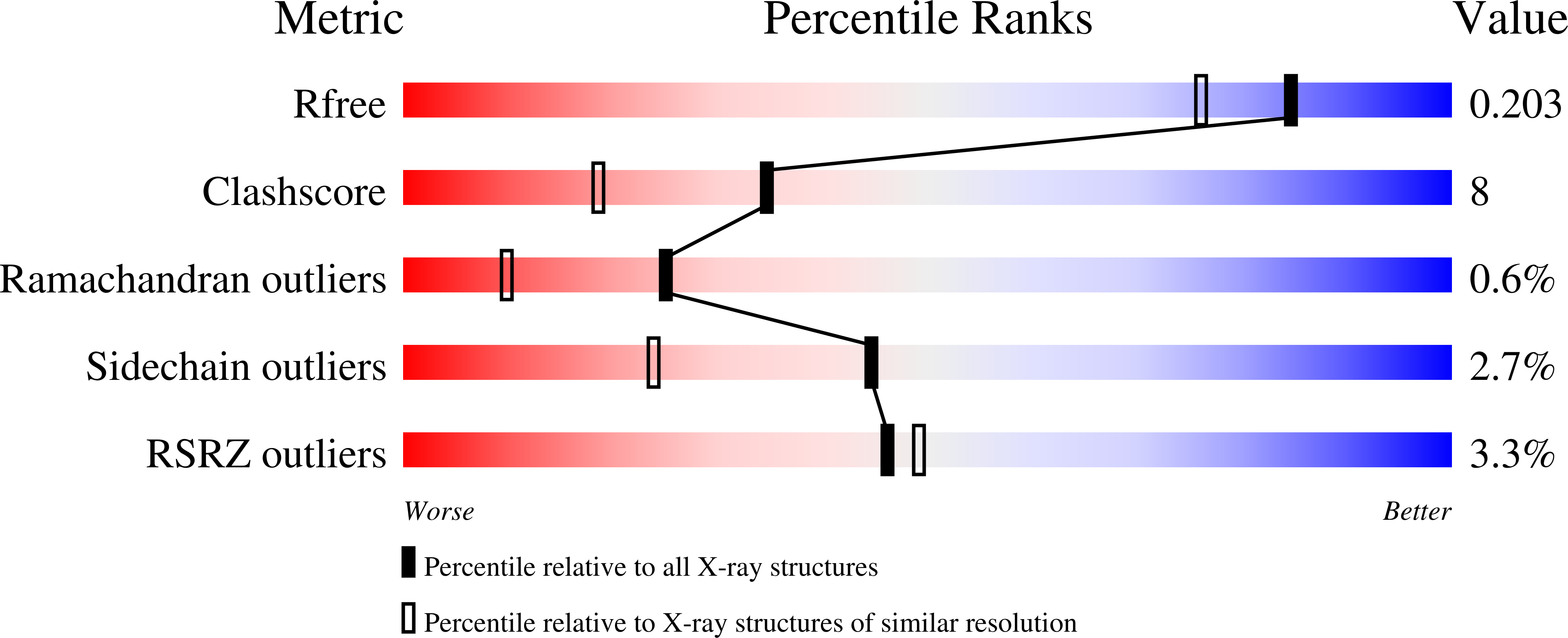
The three-dimensional crystal structure of the PduO-type corrinoid adenosyltransferase from Lactobacillus reuteri (LrPduO) has been solved to 1.68-A resolution. The functional assignment of LrPduO as a corrinoid adenosyltransferase was confirmed by in vivo and in vitro evidence. The enzyme has an apparent Km(ATP) of 2.2 microM and Km(Cobalamin) of 0.13 microM and a kcat of 0.025 s(-1). Co-crystallization of the enzyme with Mg-ATP resulted in well-defined electron density for an N-terminal loop that had been disordered in other PduO-type enzyme structures. This newly defined N-terminal loop makes up the lower portion of the enzyme active site with the other half being contributed from an adjacent subunit. These results provide the first detailed description of the enzyme active site for a PduO-type adenosyltransferase and identify a unique ATP binding motif at the protein N terminus. The molecular architecture at the active site offers valuable new insight into the role of various residues responsible for the human disease methylmalonic aciduria.
Organizational Affiliation:
Department of Biochemistry, University of Wisconsin, Madison, Wisconsin 53706, USA.