Efficacy, pharmacokinetics, and metabolism of tetrahydroquinoline inhibitors of Plasmodium falciparum protein farnesyltransferase.
Van Voorhis, W.C., Rivas, K.L., Bendale, P., Nallan, L., Horney, C., Barrett, L.K., Bauer, K.D., Smart, B.P., Ankala, S., Hucke, O., Verlinde, C.L., Chakrabarti, D., Strickland, C., Yokoyama, K., Buckner, F.S., Hamilton, A.D., Williams, D.K., Lombardo, L.J., Floyd, D., Gelb, M.H.(2007) Antimicrob Agents Chemother 51: 3659-3671
- PubMed: 17606674
- DOI: https://doi.org/10.1128/AAC.00246-07
- Primary Citation of Related Structures:
2R2L - PubMed Abstract:
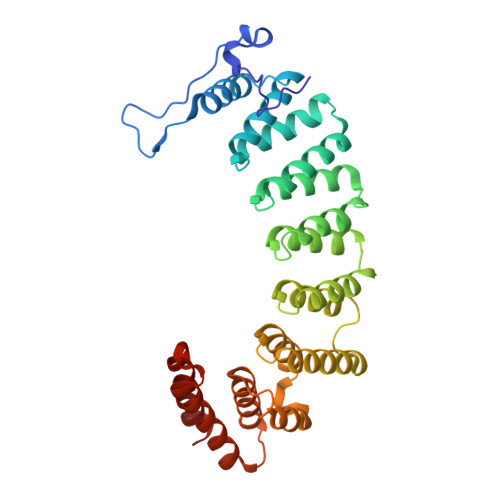
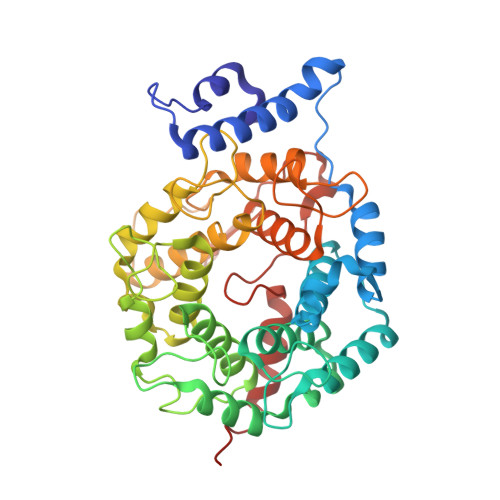
New antimalarials are urgently needed. We have shown that tetrahydroquinoline (THQ) protein farnesyltransferase (PFT) inhibitors (PFTIs) are effective against the Plasmodium falciparum PFT and are effective at killing P. falciparum in vitro. Previously described THQ PFTIs had limitations of poor oral bioavailability and rapid clearance from the circulation of rodents. In this paper, we validate both the Caco-2 cell permeability model for predicting THQ intestinal absorption and the in vitro liver microsome model for predicting THQ clearance in vivo. Incremental improvements in efficacy, oral absorption, and clearance rate were monitored by in vitro tests; and these tests were followed up with in vivo absorption, distribution, metabolism, and excretion studies. One compound, PB-93, achieved cure when it was given orally to P. berghei-infected rats every 8 h for a total of 72 h. However, PB-93 was rapidly cleared, and dosing every 12 h failed to cure the rats. Thus, the in vivo results corroborate the in vitro pharmacodynamics and demonstrate that 72 h of continuous high-level exposure to PFTIs is necessary to kill plasmodia. The metabolism of PB-93 was demonstrated by a novel technique that relied on double labeling with a radiolabel and heavy isotopes combined with radiometric liquid chromatography and mass spectrometry. The major liver microsome metabolite of PB-93 has the PFT Zn-binding N-methyl-imidazole removed; this metabolite is inactive in blocking PFT function. By solving the X-ray crystal structure of PB-93 bound to rat PFT, a model of PB-93 bound to malarial PFT was constructed. This model suggests areas of the THQ PFTIs that can be modified to retain efficacy and protect the Zn-binding N-methyl-imidazole from dealkylation.
Organizational Affiliation:
Department of Medicine, University of Washington, Room I-104-E, Health Sciences Building, Seattle, WA 98195-7185, USA. wesley@u.washington.edu