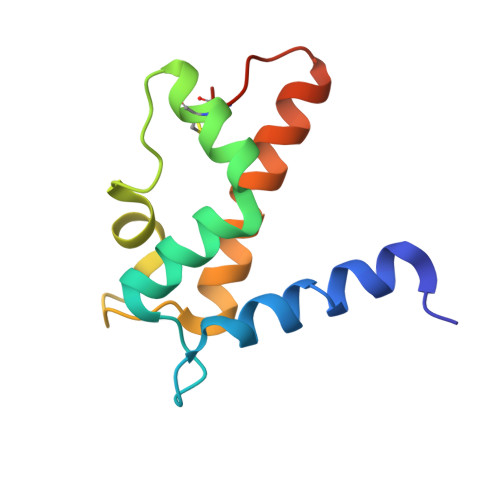
Identification and Characterization of Binding Sites on S100A7, a Participant in Cancer and Inflammation Pathways.
Leon, R., Murray, J.I., Cragg, G., Farnell, B., West, N.R., Pace, T.C., Watson, P.H., Bohne, C., Boulanger, M.J., Hof, F.(2009) Biochemistry 48: 10591
- PubMed: 19810752
- DOI: https://doi.org/10.1021/bi901330g
- Primary Citation of Related Structures:
2WOR, 2WOS - PubMed Abstract:
S100A7 (psoriasin) is a member of the S100 family of signaling proteins. It is implicated in and considered a therapeutic target for inflammation and cancer, yet no small molecule ligands for S100A7 have been identified. To begin the development of specific small molecule inhibitors of S100A7 function, we have used a series of surface binding fluorescent dyes to probe the surface hydrophobic sites. Two naphthalene-based dyes (2,6-ANS and 1,8-ANS) were found to bind S100A7 in a distinct cleft. We characterized the binding interaction by determining both the structure of S100A7 bound to 2,6-ANS and the structure of S100A7 bound to 1,8-ANS to 1.6 A. In both cases, two molecules of dye were docked such that the naphthalene groups were positioned in two symmetry-related grooves that are formed by the N-terminal helices of each monomer. We observed that Met12 acts as a gatekeeper to the binding cleft, adopting an "open" conformation for the more elongated 2,6-ANS while remaining in a "closed" conformation for the more compact 1,8-ANS. Steady-state fluorescence experiments revealed that S100A7 binds two copies of 2,6-ANS, each with a K(d) of 125 muM. Time-resolved fluorescence lifetime measurements indicated that the two molecules of 2,6-ANS bind in two independent binding sites with different fluorescence lifetimes, suggesting that the S100A7 homodimer is not perfectly symmetric in solution. Isothermal titration calorimetry studies demonstrate that S100A7 has a higher affinity for 2,6-ANS than 1,8-ANS. Yeast two-hybrid studies were also used to probe contributions of individual residues of an S100A7 triple mutant with respect to Jab1 binding. Mutation of Leu78, which forms part of the Met12 cleft occupied by 2,6-ANS, reduced the level of Jab1 binding, suggesting a potentially important role for the Met12 hydrophobic pocket in defining a Jab1 interface. Additional Y2H studies also delineate contributions of Gln88 and in particular Asp56 that shows the most significant abrogated binding to Jab1. Collectively, these data suggest a complex interaction between S100A7 and the much larger Jab1. These studies form the basis for the development of small molecule reporters and modifiers of S100A7 form and function.
Organizational Affiliation:
Department of Chemistry, University of Victoria, P.O. Box 3065, Victoria, British Columbia V8W 3V6, Canada.