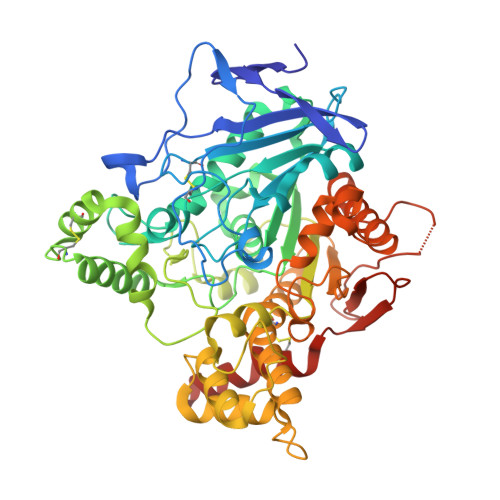
Backdoor Opening Mechanism in Acetylcholinesterase Based on X-Ray Crystallography and Molecular Dynamics Simulations.
Sanson, B., Colletier, J.P., Xu, Y., Lang, P.T., Jiang, H., Silman, I., Sussman, J.L., Weik, M.(2011) Protein Sci 20: 1114
- PubMed: 21594947
- DOI: https://doi.org/10.1002/pro.661
- Primary Citation of Related Structures:
2XI4 - PubMed Abstract:
The transient opening of a backdoor in the active-site wall of acetylcholinesterase, one of nature's most rapid enzymes, has been suggested to contribute to the efficient traffic of substrates and products. A crystal structure of Torpedo californica acetylcholinesterase in complex with the peripheral-site inhibitor aflatoxin is now presented, in which a tyrosine at the bottom of the active-site gorge rotates to create a 3.4-Å wide exit channel. Molecular dynamics simulations show that the opening can be further enlarged by movement of Trp84. The crystallographic and molecular dynamics simulation data thus point to the interface between Tyr442 and Trp84 as the key element of a backdoor, whose opening permits rapid clearance of catalysis products from the active site. Furthermore, the crystal structure presented provides a novel template for rational design of inhibitors and reactivators, including anti-Alzheimer drugs and antidotes against organophosphate poisoning.
Organizational Affiliation:
Comissariat à l'Energie Atomique, Institut de Biologie Structurale, F-38054 Grenoble, France.