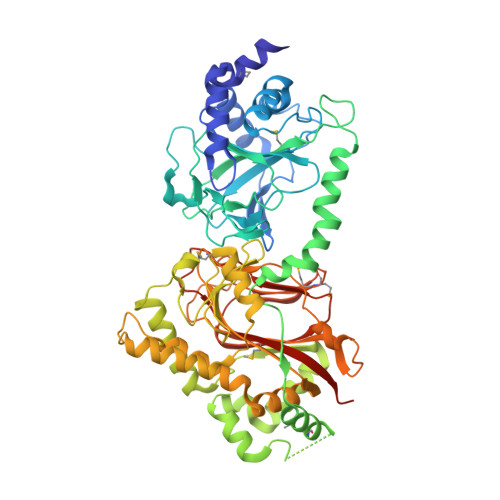
Crystal structure of Tpa1 from Saccharomyces cerevisiae, a component of the messenger ribonucleoprotein complex
Kim, H.S., Kim, H.L., Kim, K.H., Kim, D.J., Lee, S.J., Yoon, J.Y., Yoon, H.J., Lee, H.Y., Park, S.B., Kim, S.-J., Lee, J.Y., Suh, S.W.(2010) Nucleic Acids Res 38: 2099-2110
- PubMed: 20040577
- DOI: https://doi.org/10.1093/nar/gkp1151
- Primary Citation of Related Structures:
3KT1, 3KT4, 3KT7 - PubMed Abstract:
Tpa1 (for termination and polyadenylation) from Saccharomyces cerevisiae is a component of a messenger ribonucleoprotein (mRNP) complex at the 3' untranslated region of mRNAs. It comprises an N-terminal Fe(II)- and 2-oxoglutarate (2OG) dependent dioxygenase domain and a C-terminal domain. The N-terminal dioxygenase domain of a homologous Ofd1 protein from Schizosaccharomyces pombe was proposed to serve as an oxygen sensor that regulates the activity of the C-terminal degradation domain. Members of the Tpa1 family are also present in higher eukaryotes including humans. Here we report the crystal structure of S. cerevisiae Tpa1 as a representative member of the Tpa1 family. Structures have been determined as a binary complex with Fe(III) and as a ternary complex with Fe(III) and 2OG. The structures reveal that both domains of Tpa1 have the double-stranded beta-helix fold and are similar to prolyl 4-hydroxylases. However, the binding of Fe(III) and 2OG is observed in the N-terminal domain only. We also show that Tpa1 binds to poly(rA), suggesting its direct interaction with mRNA in the mRNP complex. The structural and functional data reported in this study support a role of the Tpa1 family as a hydroxylase in the mRNP complex and as an oxygen sensor.
- Department of Chemistry, College of Natural Sciences, Seoul National University, Seoul 151-742, Korea.
Organizational Affiliation: