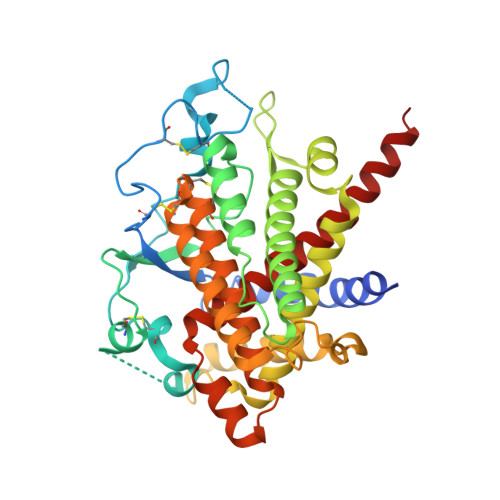
Steps in reductive activation of the disulfide-generating enzyme Ero1p
Heldman, N., Vonshak, O., Sevier, C.S., Vitu, E., Mehlman, T., Fass, D.(2010) Protein Sci 19: 1863-1876
- PubMed: 20669236
- DOI: https://doi.org/10.1002/pro.473
- Primary Citation of Related Structures:
3M31, 3NVJ - PubMed Abstract:
Ero1p is the primary catalyst of disulfide bond formation in the yeast endoplasmic reticulum (ER). Ero1p contains a pair of essential disulfide bonds that participate directly in the electron transfer pathway from substrate thiol groups to oxygen. Remarkably, elimination of certain other Ero1p disulfides by mutation enhances enzyme activity. In particular, the C150A/C295A Ero1p mutant exhibits increased thiol oxidation in vitro and in vivo and interferes with redox homeostasis in yeast cells by hyperoxidizing the ER. Inhibitory disulfides of Ero1p are thus important for enzyme regulation. To visualize the differences between de-regulated and wild-type Ero1p, we determined the crystal structure of Ero1p C150A/C295A. The structure revealed local changes compared to the wild-type enzyme around the sites of mutation, but no conformational transitions within 25 A of the active site were observed. To determine how the C150--C295 disulfide nonetheless participates in redox regulation of Ero1p, we analyzed using mass spectrometry the changes in Ero1p disulfide connectivity as a function of time after encounter with reducing substrates. We found that the C150--C295 disulfide sets a physiologically appropriate threshold for enzyme activation by guarding a key neighboring disulfide from reduction. This study illustrates the diverse and interconnected roles that disulfides can play in redox regulation of protein activity.
- Department of Structural Biology, Weizmann Institute of Science, Rehovot, Israel.
Organizational Affiliation: