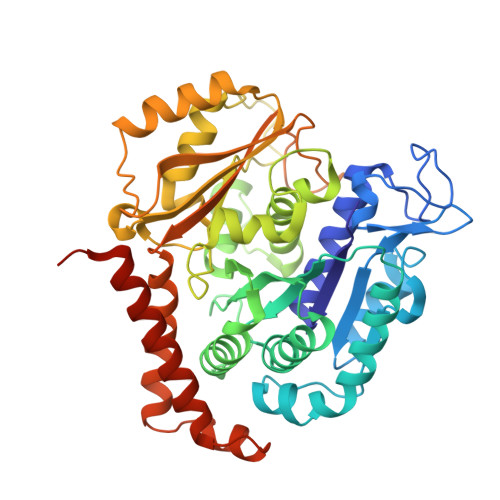
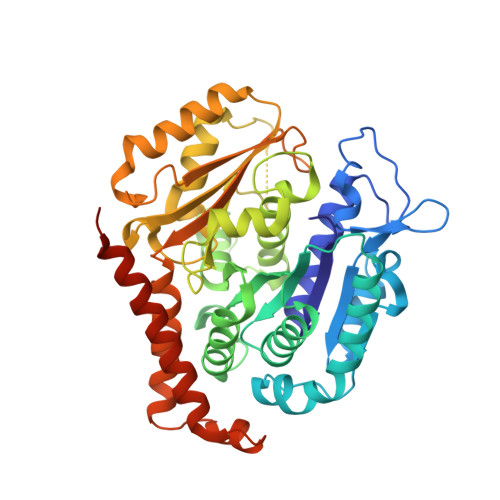
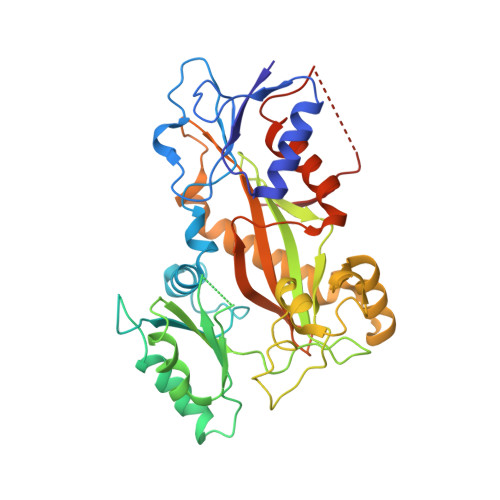
Structural basis of microtubule stabilization by laulimalide and peloruside a.
Prota, A.E., Bargsten, K., Northcote, P.T., Marsh, M., Altmann, K.H., Miller, J.H., Diaz, J.F., Steinmetz, M.O.(2014) Angew Chem Int Ed Engl 53: 1621-1625
- PubMed: 24470331
- DOI: https://doi.org/10.1002/anie.201307749
- Primary Citation of Related Structures:
4O4H, 4O4I, 4O4J, 4O4L - PubMed Abstract:
Laulimalide and peloruside A are microtubule-stabilizing agents (MSAs), the mechanism of action on microtubules of which is poorly defined. Here, using X-ray crystallography it is shown that laulimalide and peloruside A bind to a unique non-taxane site on β-tubulin and use their respective macrolide core structures to interact with a second tubulin dimer across protofilaments. At the same time, they allosterically stabilize the taxane-site M-loop that establishes lateral tubulin contacts in microtubules. Structures of ternary complexes of tubulin with laulimalide/peloruside A and epothilone A are also solved, and a crosstalk between the laulimalide/peloruside and taxane sites via the M-loop of β-tubulin is found. Together, the data define the mechanism of action of laulimalide and peloruside A on tubulin and microtubules. The data further provide a structural framework for understanding the synergy observed between two classes of MSAs in tubulin assembly and the inhibition of cancer cell growth.
- Department of Biology and Chemistry, Laboratory of Biomolecular Research, Paul Scherrer Institut, 5232 Villigen PSI (Switzerland).
Organizational Affiliation: