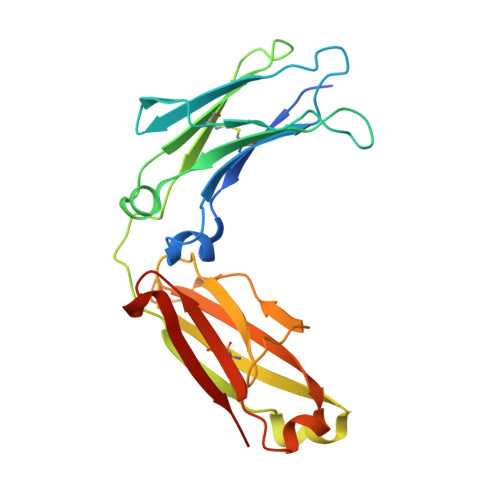
Structural characterization of anti-inflammatory immunoglobulin g fc proteins.
Ahmed, A.A., Giddens, J., Pincetic, A., Lomino, J.V., Ravetch, J.V., Wang, L.X., Bjorkman, P.J.(2014) J Mol Biology 426: 3166-3179
- PubMed: 25036289
- DOI: https://doi.org/10.1016/j.jmb.2014.07.006
- Primary Citation of Related Structures:
4Q6Y, 4Q74, 4Q7D - PubMed Abstract:
Immunoglobulin G (IgG) is a central mediator of host defense due to its ability to recognize and eliminate pathogens. The recognition and effector responses are encoded on distinct regions of IgGs. The diversity of the antigen recognition Fab domains accounts for IgG's ability to bind with high specificity to essentially any antigen. Recent studies have indicated that the Fc effector domain also displays considerable heterogeneity, accounting for its complex effector functions of inflammation, modulation, and immune suppression. Therapeutic anti-tumor antibodies, for example, require the pro-inflammatory properties of the IgG Fc to eliminate tumor cells, while the anti-inflammatory activity of intravenous IgG requires specific Fc glycans for activity. In particular, the anti-inflammatory activity of intravenous IgG is ascribed to a small population of IgGs in which the Asn297-linked complex N-glycans attached to each Fc CH2 domain include terminal α2,6-linked sialic acids. We used chemoenzymatic glycoengineering to prepare fully disialylated IgG Fc and solved its crystal structure. Comparison of the structures of asialylated Fc, sialylated Fc, and F241A Fc, a mutant that displays increased glycan sialylation, suggests that increased conformational flexibility of the CH2 domain is associated with the switch from pro-inflammatory to anti-inflammatory activity of the Fc.
- Division of Biology and Biological Engineering 114-96, California Institute of Technology, 1200 East California Boulevard, Pasadena, CA 91125, USA.
Organizational Affiliation: