Carbonic anhydrase inhibitors based on sorafenib scaffold: Design, synthesis, crystallographic investigation and effects on primary breast cancer cells.
Bozdag, M., Ferraroni, M., Ward, C., Carta, F., Bua, S., Angeli, A., Langdon, S.P., Kunkler, I.H., Al-Tamimi, A.S., Supuran, C.T.(2019) Eur J Med Chem 182: 111600-111600
- PubMed: 31419777
- DOI: https://doi.org/10.1016/j.ejmech.2019.111600
- Primary Citation of Related Structures:
6I0J, 6I0L - PubMed Abstract:
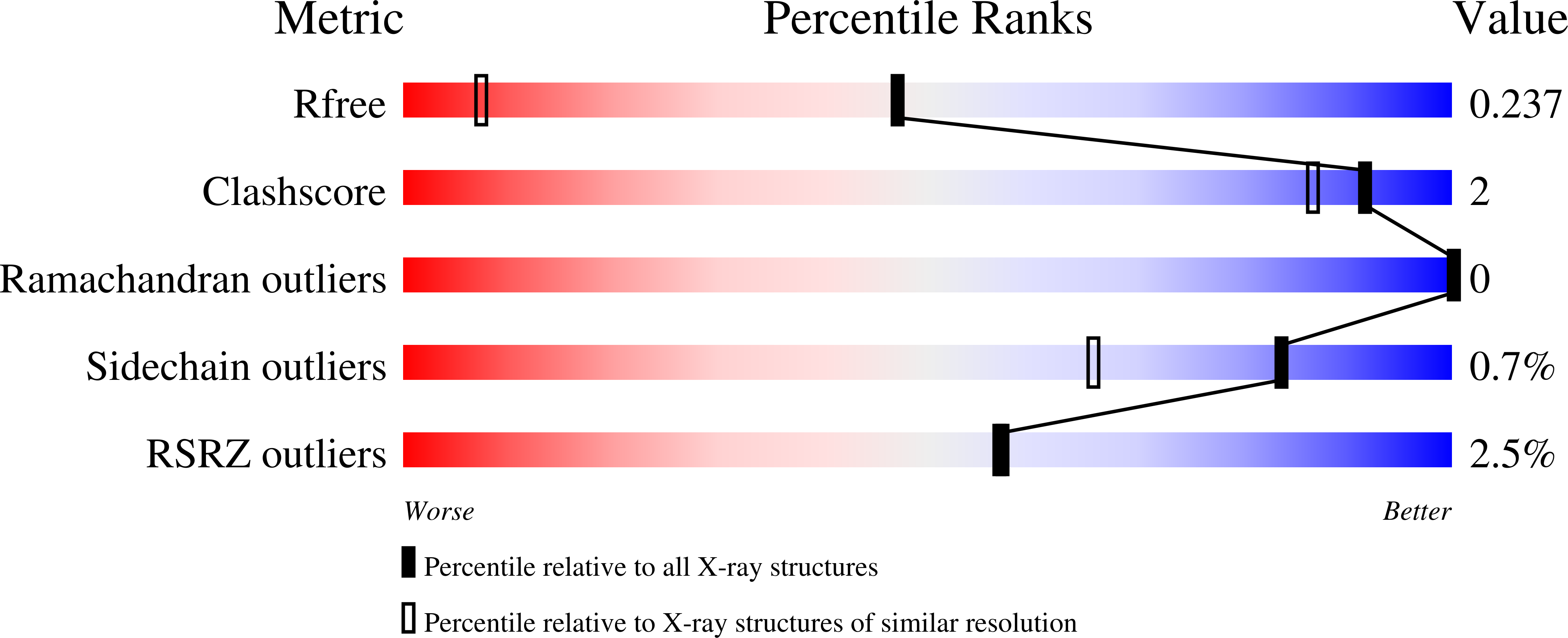
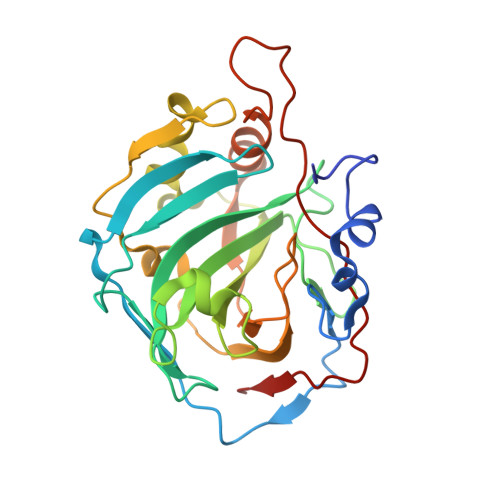
Carbonic anhydrase inhibitors (CAIs) of the sulfonamide, sulfamate and coumarin classes bearing the phenylureido tail found in the clinically used drug Sorafenib, a multikinase inhibitor actually used for the management of hepatocellular carcinomas, are reported. All compounds were assayed on human (h) CA isoforms I, II, VII and IX, involved in various pathologies. Among the sulfonamides, several compounds were selective for inhibiting hCA IX, with K I values in the low nanomolar ranges (i.e. 0.7-30.2 nM). We explored the binding modes of such compounds by means of X-ray crystallographic studies on isoform hCA I in adduct with one sulfonamide and a sulfamate inhibitor. Antiproliferative properties of some sulfamates on breast tumor cell lines were also investigated.
Organizational Affiliation:
University of Florence, NEUROFARBA Dept, Sezione di Scienze Farmaceutiche e Nutraceutiche, Via Ugo Schiff 6, 50019, Sesto Fiorentino, Florence, Italy. Electronic address: bozdag.murat@unifi.it.