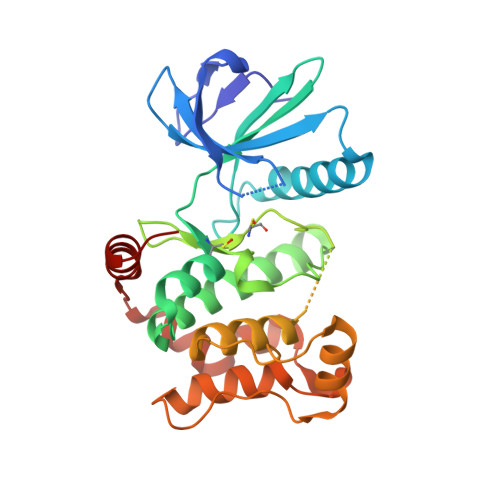
Catalytic Domain Plasticity of MKK7 Reveals Structural Mechanisms of Allosteric Activation and Diverse Targeting Opportunities.
Schroder, M., Tan, L., Wang, J., Liang, Y., Gray, N.S., Knapp, S., Chaikuad, A.(2020) Cell Chem Biol 27: 1285-1295.e4
- PubMed: 32783966
- DOI: https://doi.org/10.1016/j.chembiol.2020.07.014
- Primary Citation of Related Structures:
6YFZ, 6YG0, 6YG1, 6YG2, 6YG3, 6YG4, 6YG5, 6YG6, 6YG7, 6YZ4 - PubMed Abstract:
MKK7 (MEK7) is a key regulator of the JNK stress signaling pathway and targeting MKK7 has been proposed as a chemotherapeutic strategy. Detailed understanding of the MKK7 structure and factors that affect its activity is therefore of critical importance. Here, we present a comprehensive set of MKK7 crystal structures revealing insights into catalytic domain plasticity and the role of the N-terminal regulatory helix, conserved in all MAP2Ks, mediating kinase activation. Crystal structures harboring this regulatory helix revealed typical structural features of active kinase, providing exclusively a first model of the MAP2K active state. A small-molecule screening campaign yielded multiple scaffolds, including type II irreversible inhibitors a binding mode that has not been reported previously. We also observed an unprecedented allosteric pocket located in the N-terminal lobe for the approved drug ibrutinib. Collectively, our structural and functional data expand and provide alternative targeting strategies for this important MAP2K kinase.
- Institute of Pharmaceutical Chemistry, Goethe-University Frankfurt, Frankfurt 60438, Germany; Structural Genomics Consortium, BMLS, Goethe-University Frankfurt, Frankfurt 60438, Germany.
Organizational Affiliation: