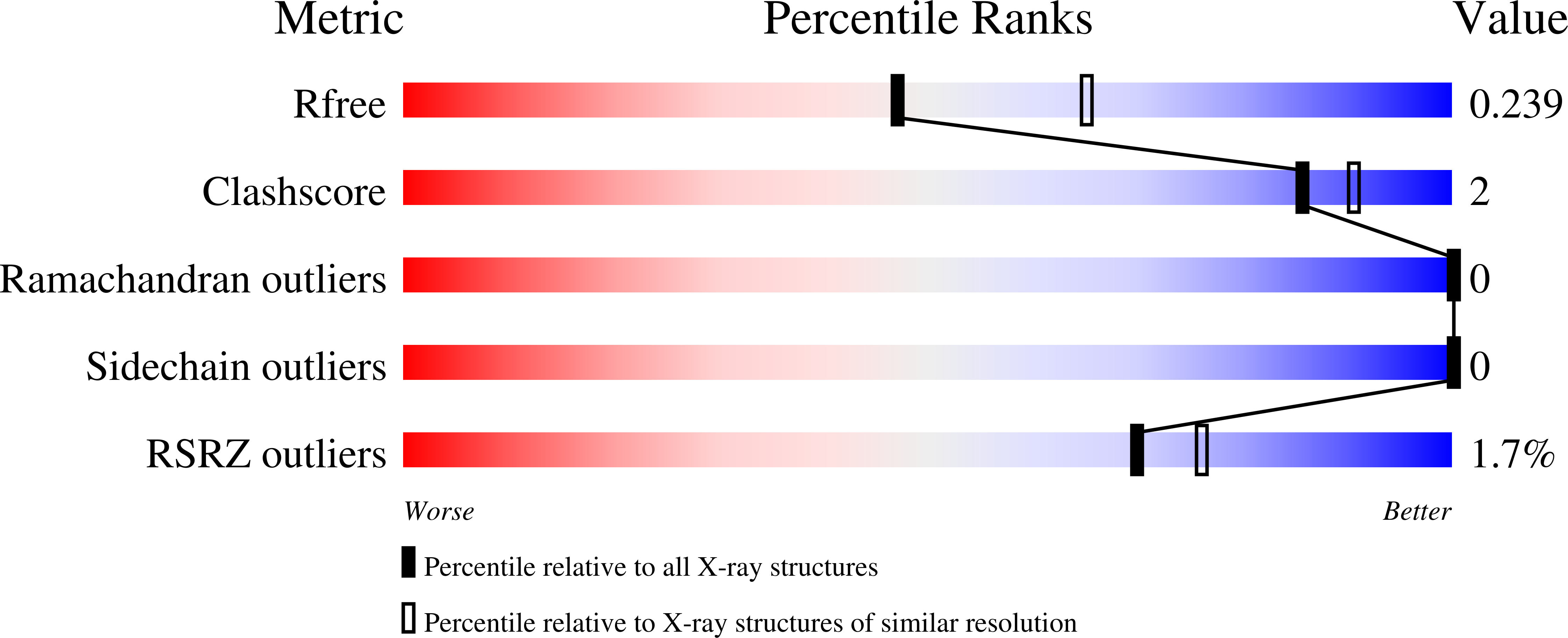
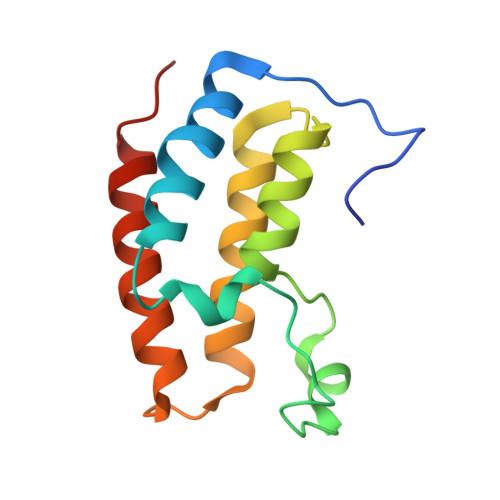
Differential BET Bromodomain Inhibition by Dihydropteridinone and Pyrimidodiazepinone Kinase Inhibitors.
Karim, R.M., Bikowitz, M.J., Chan, A., Zhu, J.Y., Grassie, D., Becker, A., Berndt, N., Gunawan, S., Lawrence, N.J., Schonbrunn, E.(2021) J Med Chem 64: 15772-15786
- PubMed: 34710325
- DOI: https://doi.org/10.1021/acs.jmedchem.1c01096
- Primary Citation of Related Structures:
5V67, 5VBO, 5VBP, 5VBQ, 5VBR, 7BJY, 7K6G, 7K6H, 7KO0, 7L6D, 7L72, 7L73, 7L9G, 7L9J, 7L9K, 7L9L, 7LAH, 7LAI, 7LAJ, 7LAK, 7LAU, 7LAY, 7LAZ, 7LB4, 7LBT, 7LEJ, 7LEK, 7LEL, 7LEM - PubMed Abstract:
BRD4 and other members of the bromodomain and extraterminal (BET) family of proteins are promising epigenetic targets for the development of novel therapeutics. Among the reported BRD4 inhibitors are dihydropteridinones and benzopyrimidodiazepinones originally designed to target the kinases PLK1, ERK5, and LRRK2. While these kinase inhibitors were identified as BRD4 inhibitors, little is known about their binding potential and structural details of interaction with the other BET bromodomains. We comprehensively characterized a series of known and newly identified dual BRD4-kinase inhibitors against all eight individual BET bromodomains. A detailed analysis of 23 novel cocrystal structures of BET-kinase inhibitor complexes in combination with direct binding assays and cell signaling studies revealed significant differences in molecular shape complementarity and inhibitory potential. Collectively, the data offer new insights into the action of kinase inhibitors across BET bromodomains, which may aid the development of drugs to inhibit certain BET proteins and kinases differentially.
Organizational Affiliation:
Drug Discovery Department, Moffitt Cancer Center, Tampa, Florida 33612, United States.