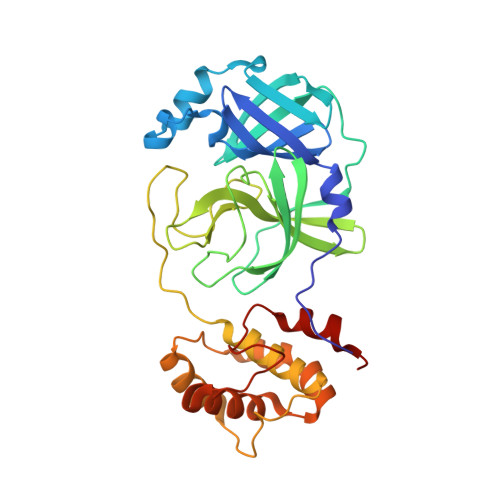
Crystal Structures of Inhibitor-Bound Main Protease from Delta- and Gamma-Coronaviruses.
Zvornicanin, S.N., Shaqra, A.M., Huang, Q.J., Ornelas, E., Moghe, M., Knapp, M., Moquin, S., Dovala, D., Schiffer, C.A., Kurt Yilmaz, N.(2023) Viruses 15
- PubMed: 36992489
- DOI: https://doi.org/10.3390/v15030781
- Primary Citation of Related Structures:
8DSU, 8E7C, 8E7N, 8FWX - PubMed Abstract:
With the spread of SARS-CoV-2 throughout the globe causing the COVID-19 pandemic, the threat of zoonotic transmissions of coronaviruses (CoV) has become even more evident. As human infections have been caused by alpha- and beta-CoVs, structural characterization and inhibitor design mostly focused on these two genera. However, viruses from the delta and gamma genera also infect mammals and pose a potential zoonotic transmission threat. Here, we determined the inhibitor-bound crystal structures of the main protease (M pro ) from the delta-CoV porcine HKU15 and gamma-CoV SW1 from the beluga whale. A comparison with the apo structure of SW1 M pro , which is also presented here, enabled the identification of structural arrangements upon inhibitor binding at the active site. The cocrystal structures reveal binding modes and interactions of two covalent inhibitors, PF-00835231 (active form of lufotrelvir) bound to HKU15, and GC376 bound to SW1 M pro . These structures may be leveraged to target diverse coronaviruses and toward the structure-based design of pan-CoV inhibitors.
Organizational Affiliation:
Department of Biochemistry and Molecular Biotechnology, University of Massachusetts Chan Medical School, Worcester, MA 01605, USA.