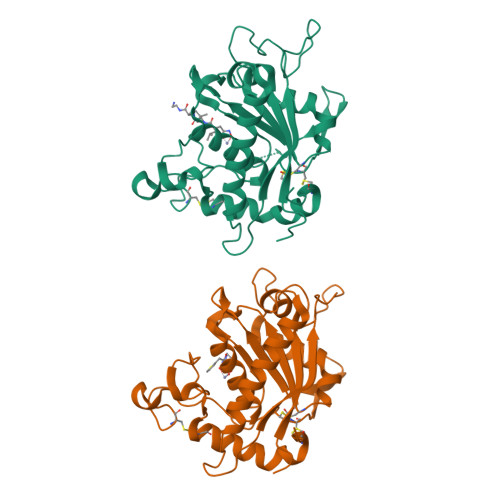
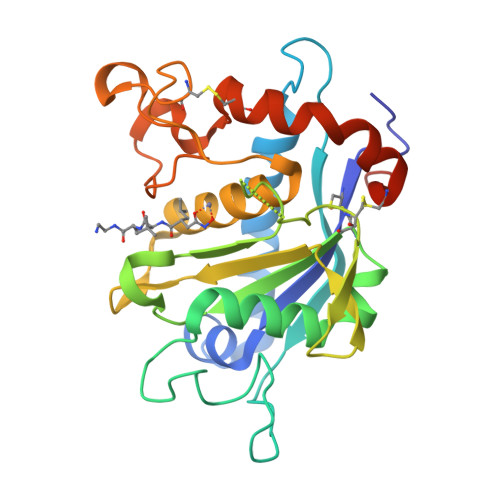
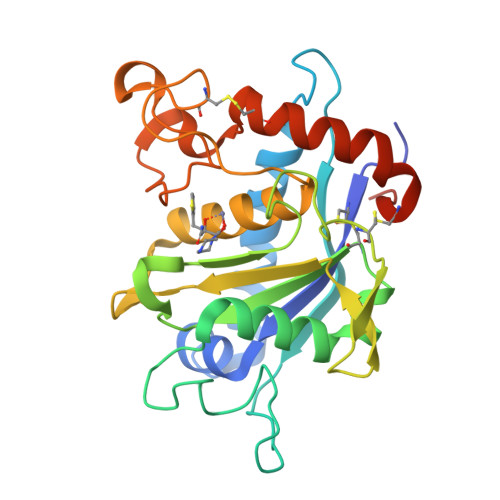
The discovery of novel tartrate-based TNF-alpha converting enzyme (TACE) inhibitors.
Rosner, K.E., Guo, Z., Orth, P., Shipps, G.W., Belanger, D.B., Chan, T.Y., Curran, P.J., Dai, C., Deng, Y., Girijavallabhan, V.M., Hong, L., Lavey, B.J., Lee, J.F., Li, D., Liu, Z., Popovici-Muller, J., Ting, P.C., Vaccaro, H., Wang, L., Wang, T., Yu, W., Zhou, G., Niu, X., Sun, J., Kozlowski, J.A., Lundell, D.J., Madison, V., McKittrick, B., Piwinski, J.J., Shih, N.Y., Arshad Siddiqui, M., Strickland, C.O.(2009) Bioorg Med Chem Lett 20: 1189-1193
- PubMed: 20022498
- DOI: https://doi.org/10.1016/j.bmcl.2009.12.004
- Primary Citation of Related Structures:
3KMC, 3KME - PubMed Abstract:
A novel series of TNF-alpha convertase (TACE) inhibitors which are non-hydroxamate have been discovered. These compounds are bis-amides of L-tartaric acid (tartrate) and coordinate to the active site zinc in a tridentate manner. They are selective for TACE over other MMP's. We report the first X-ray crystal structure for a tartrate-based TACE inhibitor.
Organizational Affiliation:
Department of Medicinal Chemistry, Schering-Plough Research Institute Cambridge, 320 Bent Street, Cambridge, MA 02141, United States.