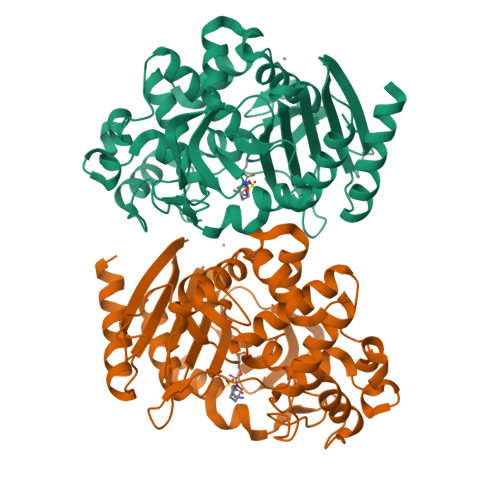
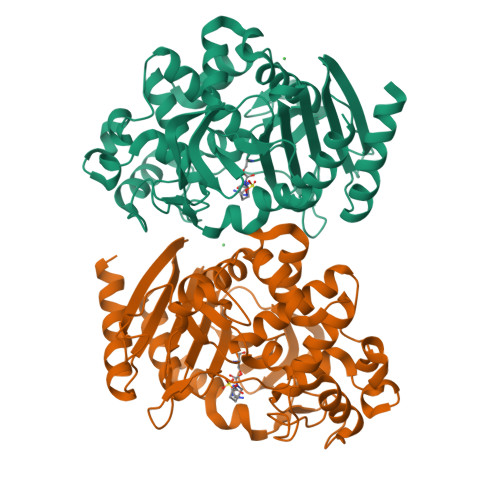
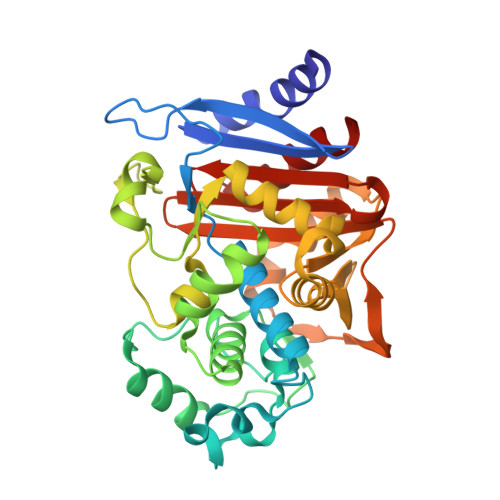
OP0595, a new diazabicyclooctane: mode of action as a serine beta-lactamase inhibitor, antibiotic and beta-lactam 'enhancer'
Morinaka, A., Tsutsumi, Y., Yamada, M., Suzuki, K., Watanabe, T., Abe, T., Furuuchi, T., Inamura, S., Sakamaki, Y., Mitsuhashi, N., Ida, T., Livermore, D.M.(2015) J Antimicrob Chemother 70: 2779-2786
- PubMed: 26089439
- DOI: https://doi.org/10.1093/jac/dkv166
- Primary Citation of Related Structures:
4X68, 4X69 - PubMed Abstract:
The production of a growing diversity of β-lactamases by Gram-negative bacteria challenges antimicrobial chemotherapy. OP0595, discovered separately by each of Meiji Seika Pharma and Fedora Pharmaceuticals, is a new diazabicyclooctane serine β-lactamase inhibitor that also acts as an antibiotic and as a β-lactamase-independent β-lactam 'enhancer'. Inhibitory activity against serine β-lactamases and affinity for PBPs were determined using nitrocefin and Bocillin FL, respectively. MICs alone and in combination with β-lactam agents were measured according to CLSI recommendations. Morphological changes in Escherichia coli were examined by phase-contrast microscopy. IC50s of OP0595 for class A and C β-lactamases were <1000 nM, with covalent binding demonstrated to the active-site serine of CTX-M-44 and AmpC enzymes. OP0595 also had direct antibiotic activity against many Enterobacteriaceae, associated with inhibition of PBP2 and conversion of the bacteria into spherical forms. Synergy between OP0595 and β-lactam agents was seen against strains producing class A and C β-lactamases vulnerable to inhibition. Lastly, OP0595 lowered the MICs of PBP3-targeted partner β-lactam agents for a non-β-lactamase-producing E. coli mutant that was resistant to OP0595 itself, indicating β-lactamase-independent 'enhancer'-based synergy. OP0595 acts in three ways: (i) as an inhibitor of class A and C β-lactamases, covalently binding at their active sites; (ii) as an antibacterial, by inhibiting PBP2 of several Enterobacteriaceae; and (iii) as an 'enhancer' of β-lactam agents that bind to other PBPs besides PBP2 for several Enterobacteriaceae. OP0595 has considerable potential to overcome resistance when it is combined with various β-lactam agents.
Organizational Affiliation:
Meiji Seika Pharma Co., Ltd, Yokohama, Japan akihiro.morinaka@meiji.com.