Structure-function analysis reveals Trichoderma virens Tsp1 to be a novel fungal effector protein modulating plant defence.
Gupta, G.D., Bansal, R., Mistry, H., Pandey, B., Mukherjee, P.K.(2021) Int J Biol Macromol 191: 267-276
- PubMed: 34547313
- DOI: https://doi.org/10.1016/j.ijbiomac.2021.09.085
- Primary Citation of Related Structures:
7CWJ, 7ESW - PubMed Abstract:
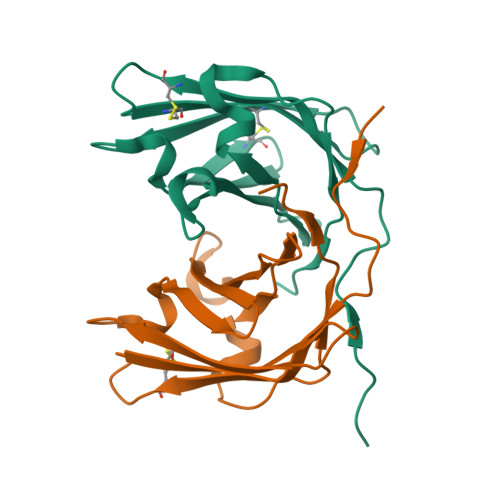
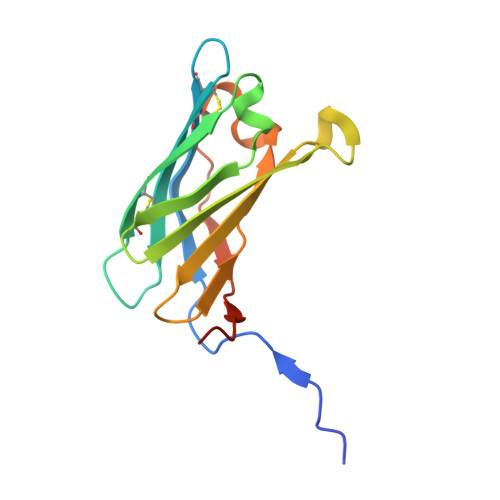
Trichoderma virens colonizes roots and develops a symbiotic relationship with plants where the fungal partner derives nutrients from plants and offers defence, in return. Tsp1, a small secreted cysteine-rich protein, was earlier found to be upregulated in co-cultivation of T. virens with maize roots. Tsp1 is well conserved in Ascomycota division of fungi, but none of its homologs have been studied yet. We have expressed and purified recombinant Tsp1, and resolved its structure to 1.25 Å resolutions, from two crystal forms, using Se-SAD methods. The Tsp1 adopts a β barrel fold and forms dimer in structure as well as in solution form. DALI based structure analysis revealed the structure similarity with two known fungal effector proteins: Alt a1 and PevD1. Structure and evolutionary analysis suggested that Tsp1 belongs to a novel effector protein family. Tsp1 acted as an inducer of salicylic acid mediated susceptibility in plants, rendering maize plants more susceptible to a necrotrophic pathogen Cochliobolus heterostrophus, as observed using plant defence assay and RT-qPCR analysis.
Organizational Affiliation:
Radiation Biology & Health Sciences Division, Bhabha Atomic Research Centre, Mumbai, India; Homi Bhabha National Institute, Training School Complex, Anushaktinagar, Mumbai, India. Electronic address: gagan@barc.gov.in.