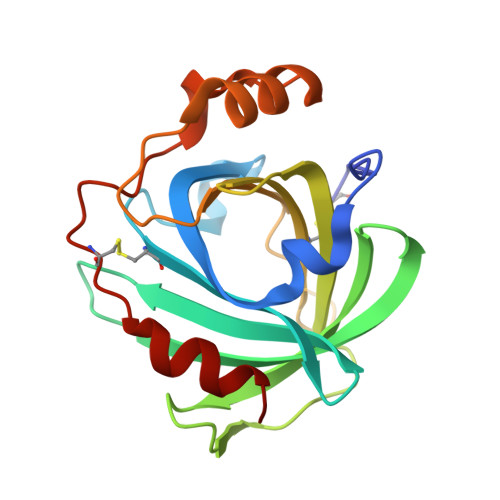
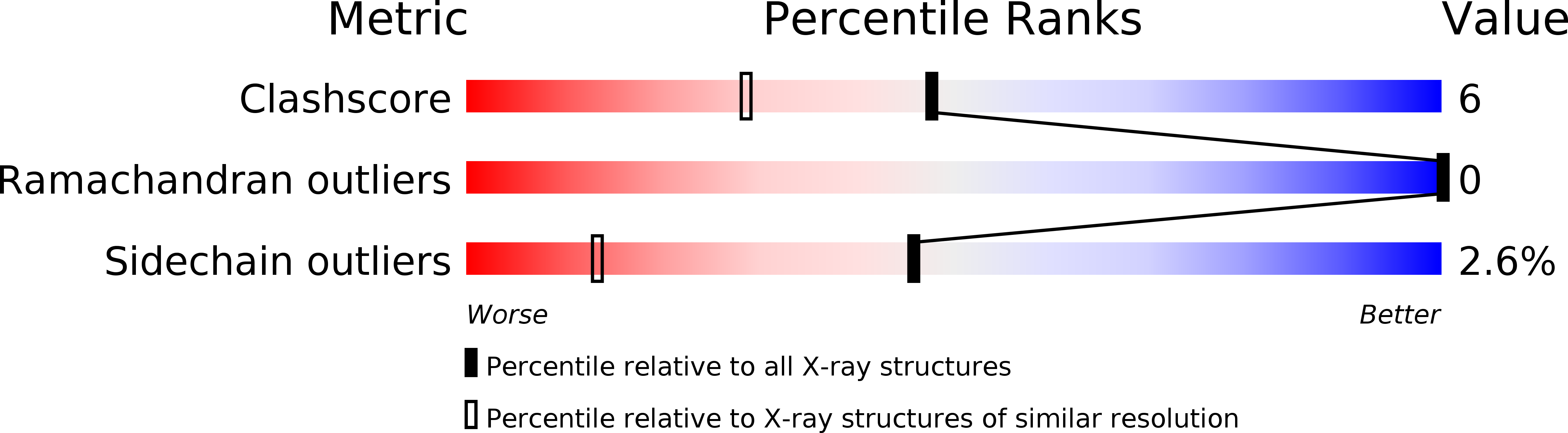Nitric oxide binding to nitrophorin 4 induces complete distal pocket burial.
Weichsel, A., Andersen, J.F., Roberts, S.A., Montfort, W.R.(2000) Nat Struct Biol 7: 551-554
- PubMed: 10876239
- DOI: https://doi.org/10.1038/76769
- Primary Citation of Related Structures:
1D3S, 1EQD, 1ERX - PubMed Abstract:
The nitrophorins comprise an unusual family of proteins that use ferric (Fe(III)) heme to transport highly reactive nitric oxide (NO) from the salivary gland of a blood sucking bug to the victim, resulting in vasodilation and reduced blood coagulation. We have determined structures of nitrophorin 4 in complexes with H2O, cyanide and nitric oxide. These structures reveal a remarkable feature: the nitrophorins have a broadly open distal pocket in the absence of NO, but upon NO binding, three or more water molecules are expelled and two loops fold into the distal pocket, resulting in the packing of hydrophobic groups around the NO molecule and increased distortion of the heme. In this way, the protein apparently forms a 'hydrophobic trap' for the NO molecule. The structures are very accurate, ranging between 1.6 and 1.4 A resolutions.
Organizational Affiliation:
Department of Biochemistry, University of Arizona, Tucson, Arizona 85721, USA.