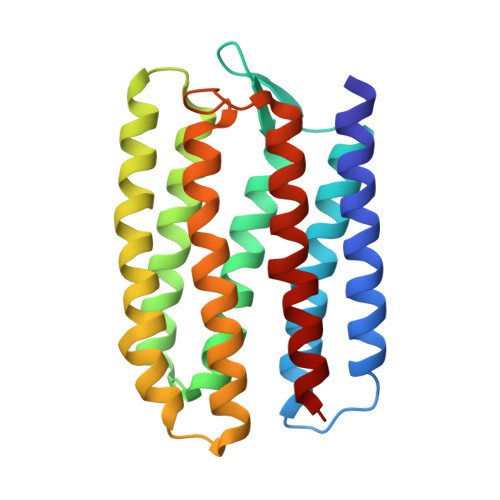
Crystal structure of sensory rhodopsin II at 2.4 angstroms: insights into color tuning and transducer interaction.
Luecke, H., Schobert, B., Lanyi, J.K., Spudich, E.N., Spudich, J.L.(2001) Science 293: 1499-1503
- PubMed: 11452084
- DOI: https://doi.org/10.1126/science.1062977
- Primary Citation of Related Structures:
1JGJ - PubMed Abstract:
We report an atomic-resolution structure for a sensory member of the microbial rhodopsin family, the phototaxis receptor sensory rhodopsin II (NpSRII), which mediates blue-light avoidance by the haloarchaeon Natronobacterium pharaonis. The 2.4 angstrom structure reveals features responsible for the 70- to 80-nanometer blue shift of its absorption maximum relative to those of haloarchaeal transport rhodopsins, as well as structural differences due to its sensory, as opposed to transport, function. Multiple factors appear to account for the spectral tuning difference with respect to bacteriorhodopsin: (i) repositioning of the guanidinium group of arginine 72, a residue that interacts with the counterion to the retinylidene protonated Schiff base; (ii) rearrangement of the protein near the retinal ring; and (iii) changes in tilt and slant of the retinal polyene chain. Inspection of the surface topography reveals an exposed polar residue, tyrosine 199, not present in bacteriorhodopsin, in the middle of the membrane bilayer. We propose that this residue interacts with the adjacent helices of the cognate NpSRII transducer NpHtrII.
Organizational Affiliation:
Department of Molecular Biology and Biochemistry, University of California, Irvine, CA 92697, USA. hudel@uci.edu