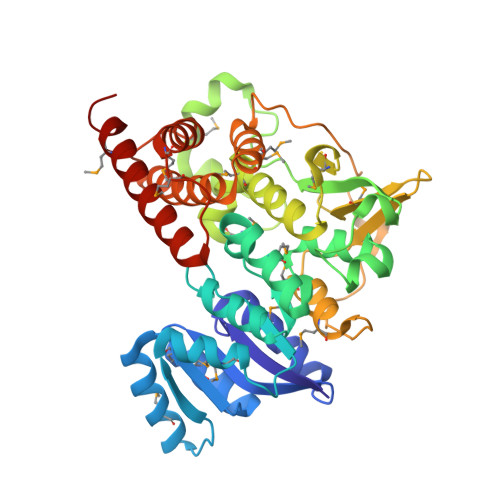
Crystal structure of 1-deoxy-D-xylulose 5-phosphate reductoisomerase complexed with cofactors: implications of a flexible loop movement upon substrate binding.
Yajima, S., Nonaka, T., Kuzuyama, T., Seto, H., Ohsawa, K.(2002) J Biochem 131: 313-317
- PubMed: 11872159
- DOI: https://doi.org/10.1093/oxfordjournals.jbchem.a003105
- Primary Citation of Related Structures:
1JVS - PubMed Abstract:
The key enzyme in the nonmevalonate pathway, 1-deoxy-D-xylulose 5-phosphate reductoisomerase (DXR), has been shown to be an effective target of antimalarial drugs. Here we report the crystal structure of DXR complexed with NADPH and a sulfate ion from Escherichia coli at 2.2 A resolution. The structure showed the presence of an extra domain, which is absent from other NADPH-dependent oxidoreductases, in addition to the conformation of catalytic residues and the substrate binding site. A flexible loop covering the substrate binding site plays an important role in the enzymatic reaction and the determination of substrate specificity.
Organizational Affiliation:
Department of Bioscience, Tokyo University of Agriculture, Setagaya-ku, Tokyo 156-8502, Japan. yshun@nodai.ac.jp