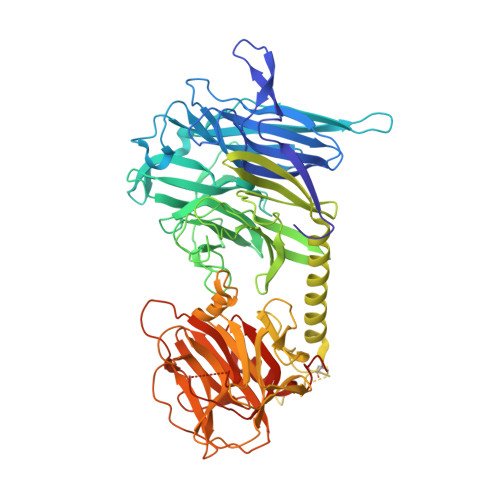
Structural basis of sialyltransferase activity in trypanosomal sialidases
Buschiazzo, A., Tavares, G.A., Campetella, O., Spinelli, S., Cremona, M.L., Paris, G., Amaya, M.F., Frasch, A.C.C., Alzari, P.M.(2000) EMBO J 19: 16-24
- PubMed: 10619840
- DOI: https://doi.org/10.1093/emboj/19.1.16
- Primary Citation of Related Structures:
1MZ5, 1MZ6 - PubMed Abstract:
The intracellular parasite Trypanosoma cruzi, the etiological agent of Chagas disease, sheds a developmentally regulated surface trans-sialidase, which is involved in key aspects of parasite-host cell interactions. Although it shares a common active site architecture with bacterial neuraminidases, the T.cruzi enzyme behaves as a highly efficient sialyltransferase. Here we report the crystal structure of the closely related Trypanosoma rangeli sialidase and its complex with inhibitor. The enzyme folds into two distinct domains: a catalytic beta-propeller fold tightly associated with a lectin-like domain. Comparison with the modeled structure of T.cruzi trans-sialidase and mutagenesis experiments allowed the identification of amino acid substitutions within the active site cleft that modulate sialyltransferase activity and suggest the presence of a distinct binding site for the acceptor carbohydrate. The structures of the Trypanosoma enzymes illustrate how a glycosidase scaffold can achieve efficient glycosyltransferase activity and provide a framework for structure-based drug design.
Organizational Affiliation:
Unité de Biochimie Structurale, Institut Pasteur, 25 rue du Dr Roux, 75724 Paris.