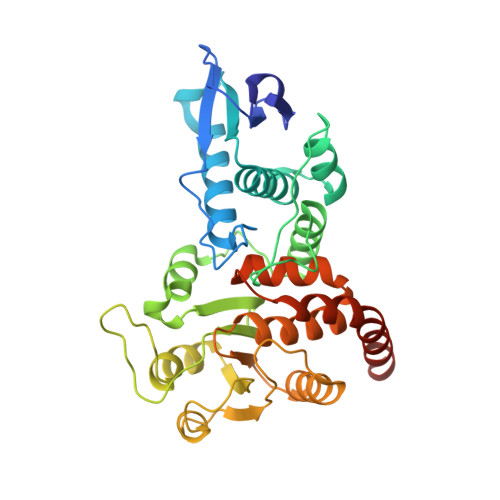
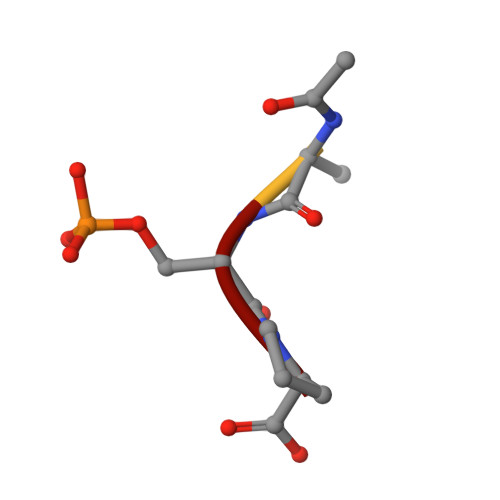
The Structure of the Cell Cycle Protein Cdc14 Reveals a Proline-Directed Protein Phosphatase
Gray, C.H., Good, V.M., Tonks, N.K., Barford, D.(2003) EMBO J 22: 3524
- PubMed: 12853468
- DOI: https://doi.org/10.1093/emboj/cdg348
- Primary Citation of Related Structures:
1OHC, 1OHD, 1OHE - PubMed Abstract:
The Cdc14 family of dual-specificity protein phosphatases (DSPs) is conserved within eukaryotes and functions to down-regulate mitotic Cdk activities, promoting cytokinesis and mitotic exit. We have integrated structural and kinetic analyses to define the molecular mechanism of the dephosphorylation reaction catalysed by Cdc14. The structure of Cdc14 illustrates a novel arrangement of two domains, each with a DSP-like fold, arranged in tandem. The C-terminal domain contains the conserved PTP motif of the catalytic site, whereas the N-terminal domain, which shares no sequence similarity with other DSPs, contributes to substrate specificity, and lacks catalytic activity. The catalytic site is located at the base of a pronounced surface channel formed by the interface of the two domains, and regions of both domains interact with the phosphopeptide substrate. Specificity for a pSer-Pro motif is mediated by a hydrophobic pocket that is capable of accommodating the apolar Pro(P+1) residue of the peptide. Our structural and kinetic data support a role for Cdc14 in the preferential dephosphorylation of proteins modified by proline-directed kinases.
Organizational Affiliation:
Section of Structural Biology, Institute of Cancer Research, Chester Beatty Laboratories, 237 Fulham Road, London SW3 6JB, UK.