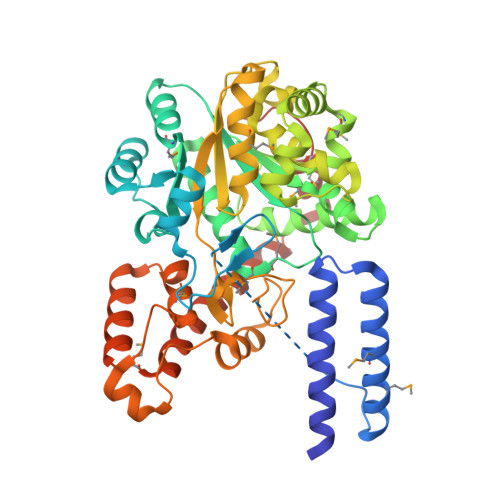
Crystal structures that suggest late development of genetic code components for differentiating aromatic side chains
Yang, X.-L., Otero, F.J., Skene, R.J., McRee, D.E., Schimmel, P., Ribas De Pouplana, L.(2003) Proc Natl Acad Sci U S A 100: 15376-15380
- PubMed: 14671330
- DOI: https://doi.org/10.1073/pnas.2136794100
- Primary Citation of Related Structures:
1Q11, 1R6T - PubMed Abstract:
Early forms of the genetic code likely generated "statistical" proteins, with similar side chains occupying the same sequence positions at different ratios. In this scenario, groups of related side chains were treated by aminoacyl-tRNA synthetases as a single molecular species until a discrimination mechanism developed that could separate them. The aromatic amino acids tryptophan, tyrosine, and phenylalanine likely constituted one of these groups. A crystal structure of human tryptophanyl-tRNA synthetase was solved at 2.1 A with a tryptophanyl-adenylate bound at the active site. A cocrystal structure of an active fragment of human tyrosyl-tRNA synthetase with its cognate amino acid analog was also solved at 1.6 A. The two structures enabled active site identifications and provided the information for structure-based sequence alignments of approximately 45 orthologs of each enzyme. Two critical positions shared by all tyrosyl-tRNA synthetases and tryptophanyl-tRNA synthetases for amino acid discrimination were identified. The variations at these two positions and phylogenetic analyses based on the structural information suggest that, in contrast to many other amino acids, discrimination of tyrosine from tryptophan occurred late in the development of the genetic code.
Organizational Affiliation:
Departments of Molecular Biology and Chemistry, The Scripps Research Institute, BCC-379, 10550 North Torrey Pines Road, La Jolla, CA 92037, USA.