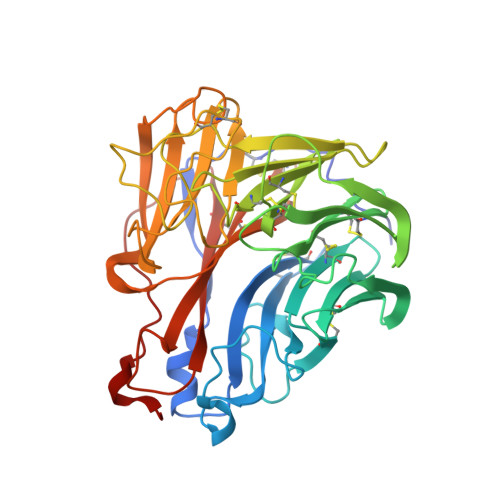
An Epidemiologically Significant Epitope of a 1998 Human Influenza Virus Neuraminidase Forms a Highly Hydrated Interface in the NA-Antibody Complex
Venkatramani, L., Bochkareva, E., Lee, J.T., Gulati, U., Laver, W.G., Bochkarev, A., Air, G.M.(2006) J Mol Biol 356: 651-663
- PubMed: 16384583
- DOI: https://doi.org/10.1016/j.jmb.2005.11.061
- Primary Citation of Related Structures:
2AEP, 2AEQ - PubMed Abstract:
The crystal structure of the complex between neuraminidase (NA) of influenza virus A/Memphis/31/98 (H3N2) and Fab of monoclonal antibody Mem5 has been determined at 2.1A resolution and shows a novel pattern of interactions compared to other NA-Fab structures. The interface buries a large area of 2400 A2 and the surfaces have high complementarity. However, the interface is also highly hydrated. There are 33 water molecules in the interface>or=95% buried from bulk solvent, but only 13 of these are isolated from other water molecules. The rest are involved in an intricate network of water-mediated hydrogen bonds throughout the interface, stabilizing the complex. Glu199 on NA, the most critical side-chain to the interaction as previously determined by escape mutant analysis and site-directed mutation, is located in a non-aqueous island. Glu199 and three other residues that contribute the major part of the antigen buried surface of the complex have mutated in human influenza viruses isolated after 1998, confirming that Mem5 identifies an epidemiologically important antigenic site. We conclude that antibody selection of NA variants is a significant component of recent antigenic drift in human H3N2 influenza viruses, supporting the idea that influenza vaccines should contain NA in addition to hemagglutinin.
Organizational Affiliation:
Department of Biochemistry and Molecular Biology, University of Oklahoma Health Sciences Center, Oklahoma City, OK 73190, USA.