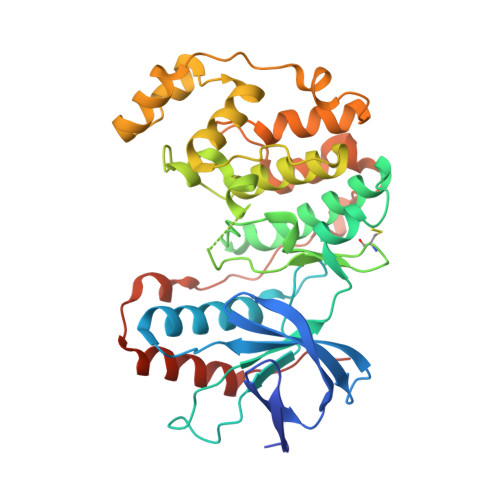
Prevention of MKK6-Dependent Activation by Binding to p38alpha MAP Kinase.
Sullivan, J.E., Holdgate, G.A., Campbell, D., Timms, D., Gerhardt, S., Breed, J., Breeze, A.L., Bermingham, A., Pauptit, R.A., Norman, R.A., Embrey, K.J., Read, J., Vanscyoc, W.S., Ward, W.H.(2005) Biochemistry 44: 16475-16490
- PubMed: 16342939
- DOI: https://doi.org/10.1021/bi051714v
- Primary Citation of Related Structures:
2BAJ, 2BAK, 2BAL, 2BAQ - PubMed Abstract:
Inhibition of p38alpha MAP kinase is a potential approach for the treatment of inflammatory disorders. MKK6-dependent phosphorylation on the activation loop of p38alpha increases its catalytic activity and affinity for ATP. An inhibitor, BIRB796, binds at a site used by the purine moiety of ATP and extends into a "selectivity pocket", which is not used by ATP. It displaces the Asp168-Phe169-Gly170 motif at the start of the activation loop, promoting a "DFG-out" conformation. Some other inhibitors bind only in the purine site, with p38alpha remaining in a "DFG-in" conformation. We now demonstrate that selectivity pocket compounds prevent MKK6-dependent activation of p38alpha in addition to inhibiting catalysis by activated p38alpha. Inhibitors using only the purine site do not prevent MKK6-dependent activation. We present kinetic analyses of seven inhibitors, whose crystal structures as complexes with p38alpha have been determined. This work includes four new crystal structures and a novel assay to measure K(d) for nonactivated p38alpha. Selectivity pocket compounds associate with p38alpha over 30-fold more slowly than purine site compounds, apparently due to low abundance of the DFG-out conformation. At concentrations that inhibit cellular production of an inflammatory cytokine, TNFalpha, selectivity pocket compounds decrease levels of phosphorylated p38alpha and beta. Stabilization of a DFG-out conformation appears to interfere with recognition of p38alpha as a substrate by MKK6. ATP competes less effectively for prevention of activation than for inhibition of catalysis. By binding to a different conformation of the enzyme, compounds that prevent activation offer an alternative approach to modulation of p38alpha.
Organizational Affiliation:
Molecular Enzymology, AstraZeneca, Mereside, Alderley Park, Macclesfield, Cheshire SK10 4TG, UK.