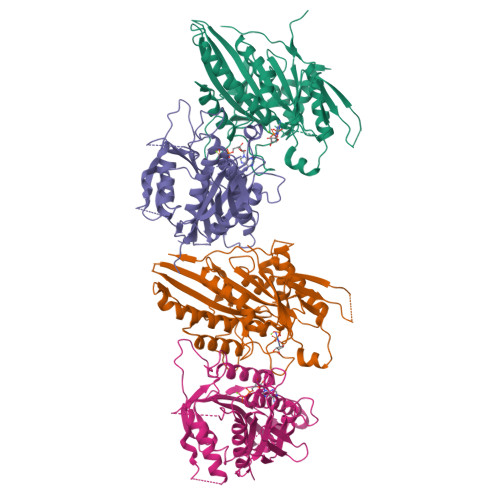
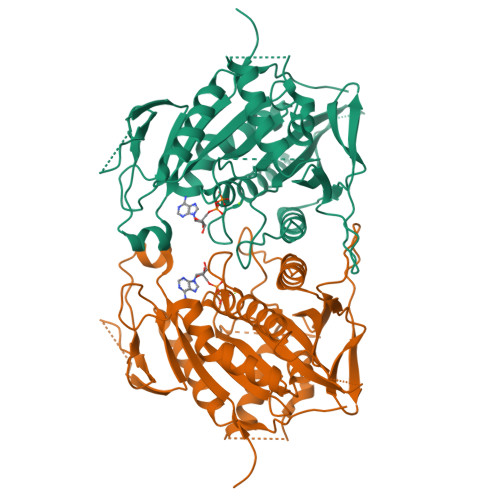
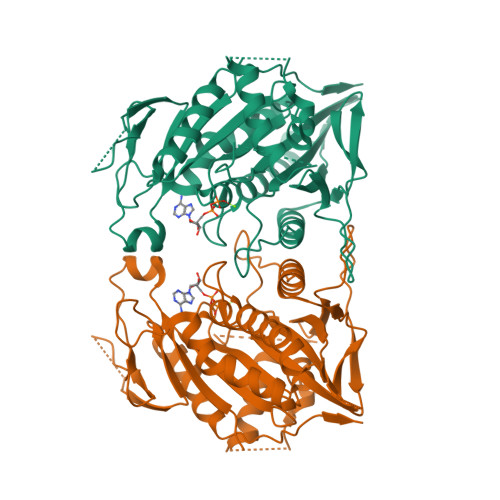
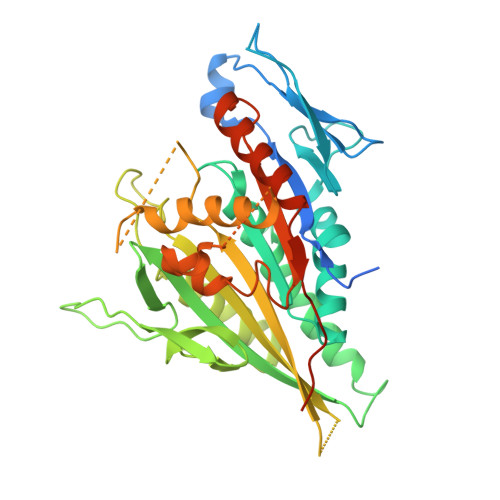
X-ray Structure and Microtubule Interaction of the Motor Domain of Neurospora crassa NcKin3, a Kinesin with Unusual Processivity
Marx, A., Muller, J., Mandelkow, E.-M., Woehlke, G., Bouchet-Marquis, C., Hoenger, A., Mandelkow, E.(2008) Biochemistry 47: 1848-1861
- PubMed: 18205396
- DOI: https://doi.org/10.1021/bi701483h
- Primary Citation of Related Structures:
2OWM - PubMed Abstract:
Neurospora crassa kinesin NcKin3 belongs to a unique fungal-specific subgroup of small Kinesin-3-related motor proteins. One of its functions appears to be the transport of mitochondria along microtubules. Here, we present the X-ray structure of a C-terminally truncated monomeric construct of NcKin3 comprising the motor domain and the neck linker, and a 3-D image reconstruction of this motor domain bound to microtubules, by cryoelectron microscopy. The protein contains Mg.ADP bound to the active site, yet the structure resembles an ATP-bound state. By comparison with structures of the Kinesin-3 motor Kif1A in different nucleotide states (Kikkawa, M. et al. (2001) Nature (London, U.K.) 411, 439-445), the NcKin3 structure corresponds to the AMPPCP complex of Kif1A rather than the AMPPNP complex. NcKin3-specific differences in the coordination of the nucleotide and asymmetric interactions between adjacent molecules in the crystal are discussed in the context of the unusual kinetics of the dimeric wild-type motor and the monomeric construct used for crystal structure analysis. The NcKin3 motor decorates microtubules at a stoichiometry of one head per alphabeta-tubulin heterodimer, thereby forming an axial periodicity of 8 nm. In spite of unusual extensions at the N-terminus and within flexible loops L2, L8a, and L12 (corresponding to the K-loop of monomeric kinesins), the microtubule binding geometry is similar to that of other members of the kinesin family.
Organizational Affiliation:
Max-Planck-Unit for Structural Molecular Biology, Notkestrasse 85, 22607 Hamburg, Germany. marx@mpasmb.desy.de