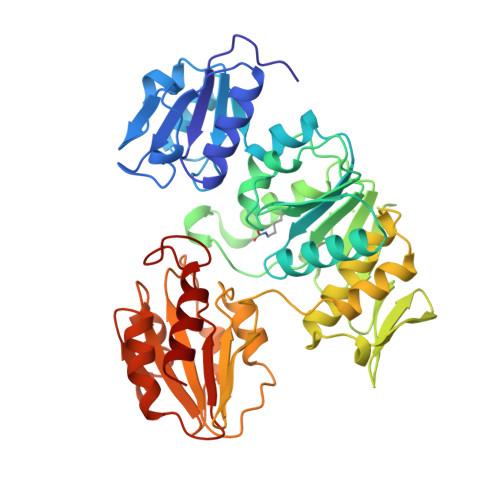
Determination of the MurD mechanism through crystallographic analysis of enzyme complexes.
Bertrand, J.A., Auger, G., Martin, L., Fanchon, E., Blanot, D., Le Beller, D., van Heijenoort, J., Dideberg, O.(1999) J Mol Biol 289: 579-590
- PubMed: 10356330
- DOI: https://doi.org/10.1006/jmbi.1999.2800
- Primary Citation of Related Structures:
2UAG, 3UAG, 4UAG - PubMed Abstract:
UDP -N- acetylmuramoyl- L -alanine: D -glutamate (MurD) ligase catalyses the addition of d -glutamate to the nucleotide precursor UDP -N- acetylmuramoyl- L -alanine (UMA). The crystal structures of three complexes of Escherichia coli MurD with a variety of substrates and products have been determined to high resolution. These include (1) the quaternary complex of MurD, the substrate UMA, the product ADP, and Mg2+, (2) the quaternary complex of MurD, the substrate UMA, the product ADP, and Mn2+, and (3) the binary complex of MurD with the product UDP - N- acetylmuramoyl- L -alanine- D -glutamate (UMAG). The reaction mechanism supported by these structures proceeds by the phosphorylation of the C-terminal carboxylate group of UMA by the gamma-phosphate group of ATP to form an acyl-phosphate intermediate, followed by the nucleophilic attack by the amino group of D-glutamate to produce UMAG. A key feature in the reaction intermediate is the presence of two magnesium ions bridging negatively charged groups.
Organizational Affiliation:
Laboratoire de Cristallographie Macromoléculaire, Institut de Biologie Structurale Jean-Pierre Ebel (CEA-CNRS), 41 rue Jules Horowitz, Grenoble Cedex 1, F-38027, France.