Alpha-Fluorophosphonates Reveal How a Phosphomutase Conserves Transition State Conformation Over Hexose Recognition in its Two-Step Reaction.
Jin, Y., Bhattasali, D., Pellegrini, E., Forget, S.M., Baxter, N.J., Cliff, M.J., Bowler, M.W., Jakeman, D.L., Blackburn, G.M., Waltho, J.P.(2014) Proc Natl Acad Sci U S A 111: 12384
- PubMed: 25104750
- DOI: https://doi.org/10.1073/pnas.1402850111
- Primary Citation of Related Structures:
2WF7, 4C4R, 4C4S, 4C4T - PubMed Abstract:
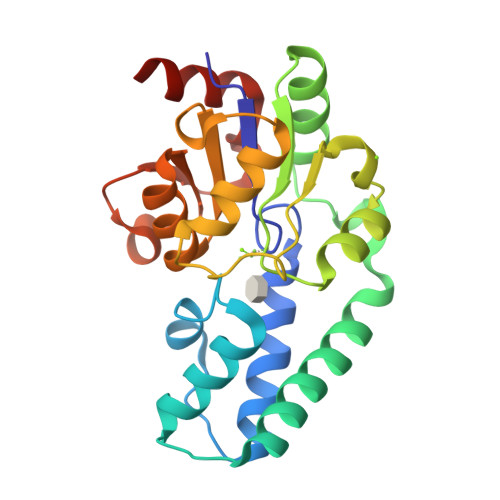
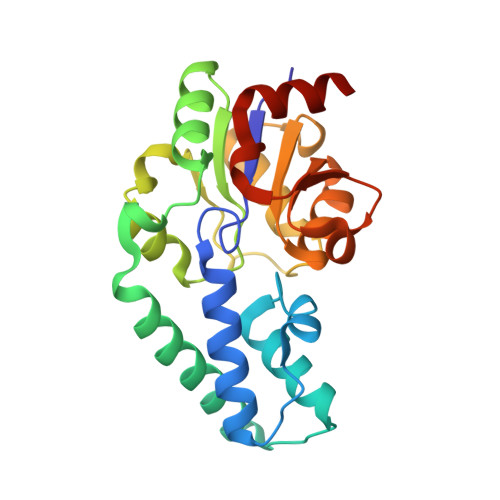
β-Phosphoglucomutase (βPGM) catalyzes isomerization of β-D-glucose 1-phosphate (βG1P) into D-glucose 6-phosphate (G6P) via sequential phosphoryl transfer steps using a β-D-glucose 1,6-bisphosphate (βG16BP) intermediate. Synthetic fluoromethylenephosphonate and methylenephosphonate analogs of βG1P deliver novel step 1 transition state analog (TSA) complexes for βPGM, incorporating trifluoromagnesate and tetrafluoroaluminate surrogates of the phosphoryl group. Within an invariant protein conformation, the β-D-glucopyranose ring in the βG1P TSA complexes (step 1) is flipped over and shifted relative to the G6P TSA complexes (step 2). Its equatorial hydroxyl groups are hydrogen-bonded directly to the enzyme rather than indirectly via water molecules as in step 2. The (C)O-P bond orientation for binding the phosphate in the inert phosphate site differs by ∼ 30° between steps 1 and 2. By contrast, the orientations for the axial O-Mg-O alignment for the TSA of the phosphoryl group in the catalytic site differ by only ∼ 5°, and the atoms representing the five phosphorus-bonded oxygens in the two transition states (TSs) are virtually superimposable. The conformation of βG16BP in step 1 does not fit into the same invariant active site for step 2 by simple positional interchange of the phosphates: the TS alignment is achieved by conformational change of the hexose rather than the protein.
Organizational Affiliation:
Department of Molecular Biology and Biotechnology, Krebs Institute, University of Sheffield, Western Bank, Sheffield S10 2TN, United Kingdom;