Crystal Structure of the C-Terminal Domain of the Salmonella Type III Secretion System Export Apparatus Protein Inva.
Worrall, L.J., Vuckovic, M., Strynadka, N.C.J.(2010) Protein Sci 19: 1091
- PubMed: 20306492
- DOI: https://doi.org/10.1002/pro.382
- Primary Citation of Related Structures:
2X49, 2X4A - PubMed Abstract:
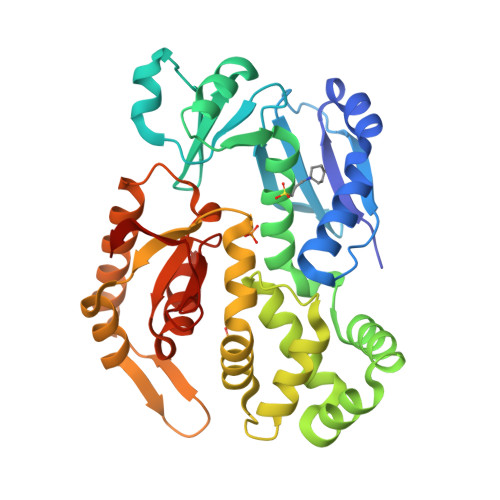
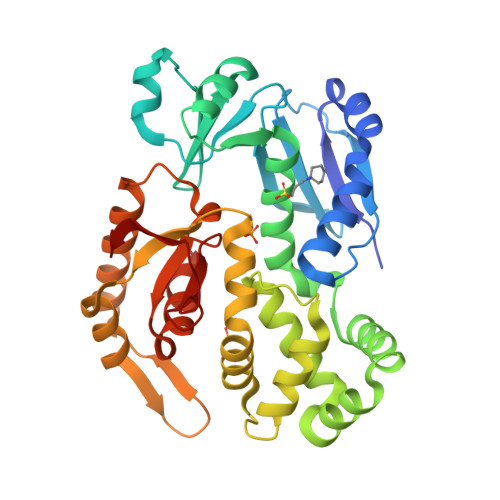
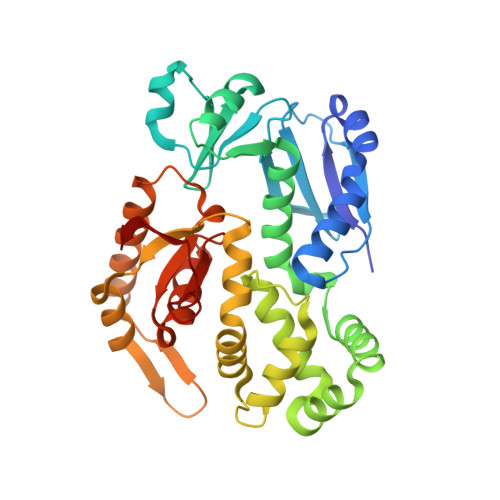
InvA is a prominent inner-membrane component of the Salmonella type III secretion system (T3SS) apparatus, which is responsible for regulating virulence protein export in pathogenic bacteria. InvA is made up of an N-terminal integral membrane domain and a C-terminal cytoplasmic domain that is proposed to form part of a docking platform for the soluble export apparatus proteins notably the T3SS ATPase InvC. Here, we report the novel crystal structure of the C-terminal domain of Salmonella InvA which shows a compact structure composed of four subdomains. The overall structure is unique although the first and second subdomains exhibit structural similarity to the peripheral stalk of the A/V-type ATPase and a ring building motif found in other T3SS proteins respectively.
Organizational Affiliation:
Centre for Blood Research, Department of Biochemistry and Molecular Biology, University of British Columbia, Life Sciences Centre, British Columbia, Canada V6T 1Z3.