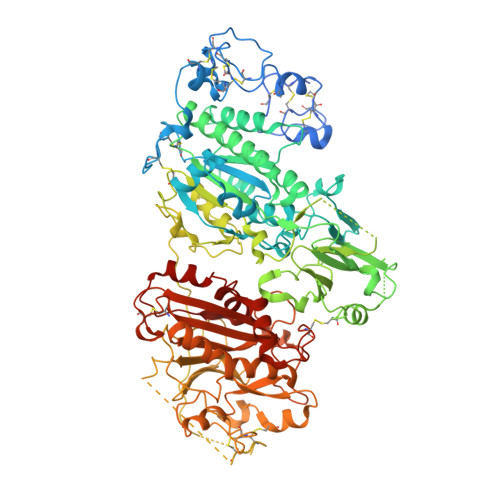
Structural Basis of Substrate Discrimination and Integrin Binding by Autotaxin.
Hausmann, J., Kamtekar, S., Christodoulou, E., Day, J.E., Wu, T., Fulkerson, Z., Albers, H.M.H.G., Van Meeteren, L.A., Houben, A.J., Van Zeijl, L., Jansen, S., Andries, M., Hall, T., Pegg, L.E., Benson, T.E., Kasiem, M., Harlos, K., Kooi, C.W., Smyth, S.S., Ovaa, H., Bollen, M., Morris, A.J., Moolenaar, W.H., Perrakis, A.(2011) Nat Struct Mol Biol 18: 198
- PubMed: 21240271
- DOI: https://doi.org/10.1038/nsmb.1980
- Primary Citation of Related Structures:
2XR9, 2XRG - PubMed Abstract:
Autotaxin (ATX, also known as ectonucleotide pyrophosphatase/phosphodiesterase-2, ENPP2) is a secreted lysophospholipase D that generates the lipid mediator lysophosphatidic acid (LPA), a mitogen and chemoattractant for many cell types. ATX-LPA signaling is involved in various pathologies including tumor progression and inflammation. However, the molecular basis of substrate recognition and catalysis by ATX and the mechanism by which it interacts with target cells are unclear. Here, we present the crystal structure of ATX, alone and in complex with a small-molecule inhibitor. We have identified a hydrophobic lipid-binding pocket and mapped key residues for catalysis and selection between nucleotide and phospholipid substrates. We have shown that ATX interacts with cell-surface integrins through its N-terminal somatomedin B-like domains, using an atypical mechanism. Our results define determinants of substrate discrimination by the ENPP family, suggest how ATX promotes localized LPA signaling and suggest new approaches for targeting ATX with small-molecule therapeutic agents.
Organizational Affiliation:
Division of Biochemistry, The Netherlands Cancer Institute, Amsterdam, The Netherlands.