Evolution of the Receptor Binding Properties of the Influenza A(H3N2) Hemagglutinin.
Lin, Y.P., Xiong, X., Wharton, S.A., Martin, S.R., Coombs, P.J., Vachieri, S.G., Christodoulou, E., Walker, P.A., Liu, J., Skehel, J.J., Gamblin, S.J., Hay, A.J., Daniels, R.S., Mccauley, J.W.(2012) Proc Natl Acad Sci U S A 109: 21474
- PubMed: 23236176
- DOI: https://doi.org/10.1073/pnas.1218841110
- Primary Citation of Related Structures:
2YP2, 2YP3, 2YP4, 2YP5, 2YP7, 2YP8, 2YP9, 2YPG - PubMed Abstract:
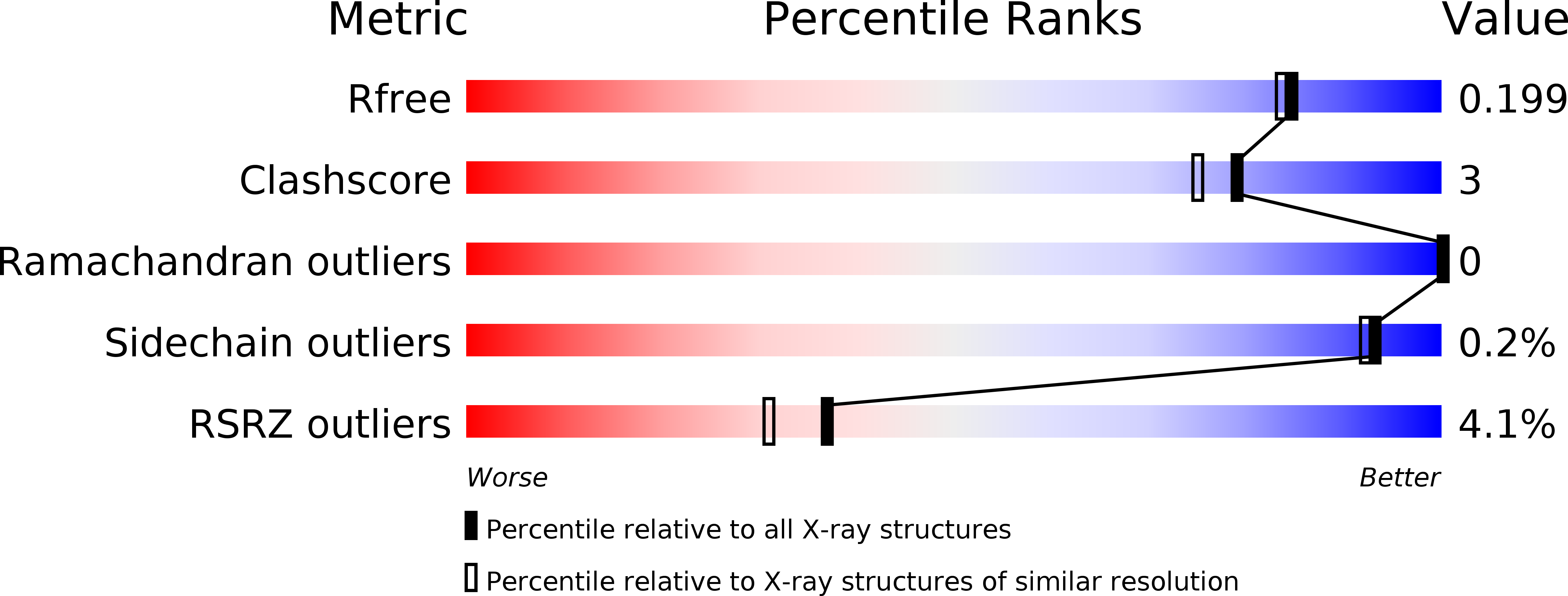
The hemagglutinin (HA) of influenza A(H3N2) virus responsible for the 1968 influenza pandemic derived from an avian virus. On introduction into humans, its receptor binding properties had changed from a preference for avian receptors (α2,3-linked sialic acid) to a preference for human receptors (α2,6-linked sialic acid). By 2001, the avidity of human H3 viruses for avian receptors had declined, and since then the affinity for human receptors has also decreased significantly. These changes in receptor binding, which correlate with increased difficulties in virus propagation in vitro and in antigenic analysis, have been assessed by virus hemagglutination of erythrocytes from different species and quantified by measuring virus binding to receptor analogs using surface biolayer interferometry. Crystal structures of HA-receptor analog complexes formed with HAs from viruses isolated in 2004 and 2005 reveal significant differences in the conformation of the 220-loop of HA1, relative to the 1968 structure, resulting in altered interactions between the HA and the receptor analog that explain the changes in receptor affinity. Site-specific mutagenesis shows the HA1 Asp-225→Asn substitution to be the key determinant of the decreased receptor binding in viruses circulating since 2005. Our results indicate that the evolution of human influenza A(H3N2) viruses since 1968 has produced a virus with a low propensity to bind human receptor analogs, and this loss of avidity correlates with the marked reduction in A(H3N2) virus disease impact in the last 10 y.
Organizational Affiliation:
Division of Virology, Medical Research Council National Institute for Medical Research, London NW7 1AA, United Kingdom.