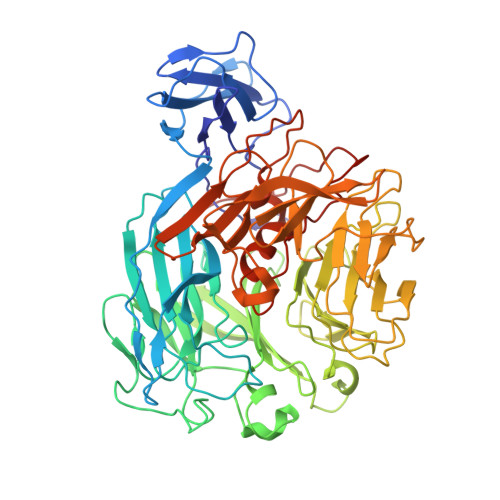
A Novel Structural Fold in Polysaccharide Lyases: BACILLUS SUBTILIS FAMILY 11 RHAMNOGALACTURONAN LYASE YesW WITH AN EIGHT-BLADED -PROPELLER
Ochiai, A., Itoh, T., Maruyama, Y., Kawamata, A., Mikami, B., Hashimoto, W., Murata, K.(2007) J Biol Chem 282: 37134-37145
- PubMed: 17947240
- DOI: https://doi.org/10.1074/jbc.M704663200
- Primary Citation of Related Structures:
2Z8R, 2Z8S - PubMed Abstract:
Rhamnogalacturonan (RG) lyase produced by plant pathogenic and saprophytic microbes plays an important role in degrading plant cell walls. An extracellular RG lyase YesW from saprophytic Bacillus subtilis is a member of polysaccharide lyase family 11 and cleaves glycoside bonds in polygalacturonan as well as RG type-I through a beta-elimination reaction. Crystal structures of YesW and its complex with galacturonan disaccharide, a reaction product analogue, were determined at 1.4 and 2.5 A resolutions with final R-factors of 16.4% and 16.6%, respectively. The enzyme is composed of an eight-bladed beta-propeller with a deep cleft in the center as a basic scaffold, and its structural fold has not been seen in polysaccharide lyases analyzed thus far. Structural analysis of the disaccharide-bound YesW and a site-directed mutagenesis study suggested that Arg-452 and Lys-535 stabilize the carboxyl group of the acidic polysaccharide molecule and Tyr-595 makes a stack interaction with the sugar pyranose ring. In addition to amino acid residues binding to the disaccharide, one calcium ion, which is coordinated by Asp-401, Glu-422, His-363, and His-399, may mediate the enzyme activity. This is, to our knowledge, the first report of a new structural category with a beta-propeller fold in polysaccharide lyases and provides structural insights into substrate binding by RG lyase.
Organizational Affiliation:
Laboratory of Basic and Applied Molecular Biotechnology, Graduate School of Agriculture, Kyoto University, Japan.