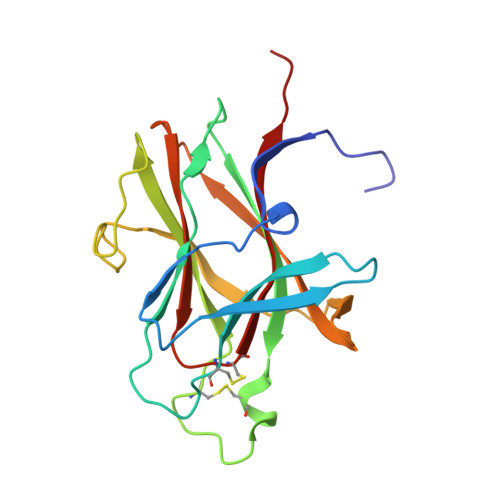
Structure of the ligand-binding domain of the EphB2 receptor at 2 A resolution.
Goldgur, Y., Paavilainen, S., Nikolov, D., Himanen, J.P.(2009) Acta Crystallogr Sect F Struct Biol Cryst Commun 65: 71-74
- PubMed: 19193989
- DOI: https://doi.org/10.1107/S1744309108043078
- Primary Citation of Related Structures:
3ETP - PubMed Abstract:
Eph tyrosine kinase receptors, the largest group of receptor tyrosine kinases, and their ephrin ligands are important mediators of cell-cell communication regulating cell attachment, shape and mobility. Recently, several Eph receptors and ephrins have also been found to play important roles in the progression of cancer. Structural and biophysical studies have established detailed information on the binding and recognition of Eph receptors and ephrins. The initial high-affinity binding of Eph receptors to ephrin occurs through the penetration of an extended G-H loop of the ligand into a hydrophobic channel on the surface of the receptor. Consequently, the G-H loop-binding channel of Eph receptors is the main target in the search for Eph antagonists that could be used in the development of anticancer drugs and several peptides have been shown to specifically bind Eph receptors and compete with the cognate ephrin ligands. However, the molecular details of the conformational changes upon Eph/ephrin binding have remained speculative, since two of the loops were unstructured in the original model of the free EphB2 structure and their conformational changes upon ligand binding could consequently not be analyzed in detail. In this study, the X-ray structure of unbound EphB2 is reported at a considerably higher 2 A resolution, the conformational changes that the important receptor loops undergo upon ligand binding are described and the consequences that these findings have for the development of Eph antagonists are discussed.
Organizational Affiliation:
Structural Biology Program, Memorial Sloan-Kettering Cancer Center, 1275 York Avenue, New York 10065, USA.