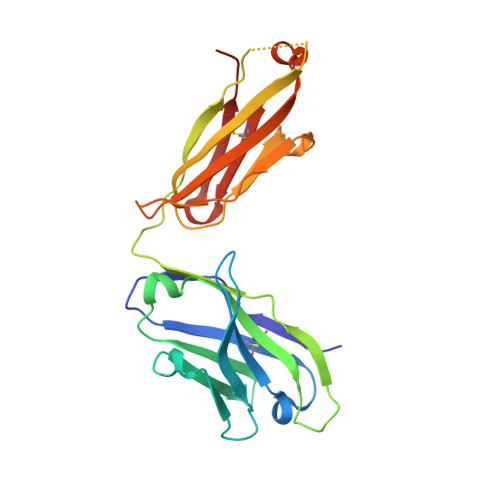
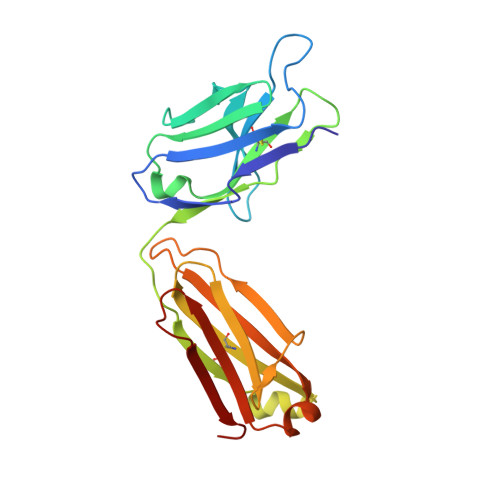
Biochemical and structural characterization of lysophosphatidic Acid binding by a humanized monoclonal antibody.
Fleming, J.K., Wojciak, J.M., Campbell, M.A., Huxford, T.(2011) J Mol Biol 408: 462-476
- PubMed: 21392506
- DOI: https://doi.org/10.1016/j.jmb.2011.02.061
- Primary Citation of Related Structures:
3QCT, 3QCU, 3QCV - PubMed Abstract:
Lysophosphatidic acid (LPA) is a common product of glycerophospholipid metabolism and an important mediator of signal transduction. Aberrantly high LPA concentrations accompany multiple disease states. One potential approach for treatment of these diseases, therefore, is the therapeutic application of antibodies that recognize and bind LPA as their antigen. We have determined the X-ray crystal structure of an anti-LPA antibody (LT3015) Fab fragment in its antigen-free form to 2.15 Å resolution and in complex with two LPA isotypes (14:0 and 18:2) to resolutions of 1.98 and 2.51 Å, respectively. The variable CDR (complementarity-determining region) loops at the antigen binding site adopt nearly identical conformations in the free and antigen-bound crystal structures. The crystallographic models reveal that the LT3015 antibody employs both heavy- and light-chain CDR loops to create a network of eight hydrogen bonds with the glycerophosphate head group of its LPA antigen. The head group is almost completely excluded from contact with solvent, while the hydrocarbon tail is partially solvent-exposed. In general, mutation of amino acid residues at the antigen binding site disrupts LPA binding. However, the introduction of particular mutations chosen strategically on the basis of the structures can positively influence LPA binding affinity. Finally, these structures elucidate the exquisite specificity demonstrated by an anti-lipid antibody for binding a structurally simple and seemingly unconstrained target molecule.
Organizational Affiliation:
Structural Biochemistry Laboratory, Department of Chemistry and Biochemistry, San Diego State University, 5500 Campanile Drive, San Diego, CA 92182-1030, USA.